In Silico Validation Studies of Cyanobacterial Bioactive Compounds Against Α-amylase and Α-glucosidase Markers in Type 2 Diabetes Mellitus
Abstract
Aim
The study aims to assess the binding efficiency of cyanobacterial compounds against key Type 2 Diabetes Mellitus (T2DM) targets, α-amylase and α-glucosidase, using an in-silico approach. Additionally, it aims to design drugs with minimal adverse effects or no toxicity to inhibit the complications and help in the management of T2DM.
Methods
Twenty-five (25) cyanobacterial bioactive compounds were sourced from various cyanobacterial strains via the PubChem database. The three-dimensional structures of the target proteins, α-amylase (1KB3) and α-glucosidase (1QOX) were obtained from RCSB PDB and visualized using Discovery Studio Visualizer 3.0. Molecular docking was performed using AutoDock 4.2 and Cygwin.
Results
Studies revealed that Ethyl tumonoate A, Debromoaplysiatoxin, and Scytoscalalrol exhibited higher binding interactions with α-amylase (1KB3), while Ambiguine I Isonitrile, Scytoscalalrol, and Cylindrospermopsin displayed higher binding affinities with α-glucosidase (1QOX) among the tested cyanobacterial bioactive compounds. These compounds exhibited greater binding affinities compared to synthetic drugs like metformin (-7.66 kcal/mol) and acarbose (-8.86 kcal/mol).
Conclusion
Our findings suggest that cyanobacterial bioactive compounds, particularly Ethyl tumonoate A, Ambiguine I Isonitrile, Cylindrospermopsin, and Scytoscalalrol, possess potential binding affinities with T2DM-related targets, making them promising lead compounds for the development of novel drugs with fewer side effects for the management of T2DM and its associated complications.
1. INTRODUCTION
A chronic condition of the metabolism of carbo- hydrates, lipids, and proteins known as diabetes mellitus (DM) is characterized by elevated postprandial and fasting blood glucose levels (hyperglycemia). Diabetes mellitus is caused by either impaired insulin action or insufficient insulin production [1]. Abnormal metabolism of lipids and proteins is a key provider to illness and death in many developing and developed countries, including China, India United States and several other countries. Diabetes mellitus has significant statistical implications. According to current data published by the World Health Organization (WHO) up to March 2013, the incidence of diabetes cases has increased from 153 million patients in 1980 to 347 million persons globally with diabetes mellitus in 2008. (DM). Furthermore, it is estimated that by 2025, this number will have risen to 380 million, accounting for 7.1 percent of the world's adult population [2] and it is supposed to be 500 million up to 2030 [3]. The selection of proper preventive measures is necessary to reduce the disease burden on the health and economy. Although several synthetic drugs exist for the treatment of diabetes mellitus, none of these medications are low-priced or completely effective. Furthermore, prolonged use of these drugs may produce undesirable side effects [4]. As a result, diabetes mellitus is a serious public well-being issue that affects millions of people in high, middle and low-income nations [5], and abnormal metabolism of proteins and lipids is a major contributor to morbidity and mortality in many developed and developing countries, including the United States and India. The insulin functions, including secretion, site of action and action, when altered or defected, are the principal factors responsible for diabetes. The severity of diabetes is connected with the consecutive destruction of microstructures, mainly the retina, nephron, and neurons, affecting eyes, kidneys and nerves. The disease is also associated with several cardiovascular diseases and related complications [6, 7]. Apart from that, oxidative stress also plays a vital role in the development of diabetes mellitus and its complications [4]. The selection of proper preventive measures is necessary to reduce the disease burden on the health and economy. One of the main salivary proteins, alpha-amylase, has been suggested as a sensitive, non-invasive biomarker whose primary function is the enzymatic digestion of carbohydrates by the hydrolysis of starch to sugar and maltose [8]. Treatment for abnormalities in carbohydrate uptake, including diabetes, obesity, dental caries, and periodontal illnesses, includes inhibition of the enzyme α-amylase, which is involved in the digestion of starch and glycogen. It serves as a marker for the diagnosis of diabetes mellitus when it is found to be significantly higher in diabetics than in non-diabetics, leading to excessive glucose production. A few studies have also suggested that reducing the absorption of complex carbs while using a salivary α-amylase inhibitor may help manage blood sugar levels [8]. Therefore, there is a need for additional research to support the earlier results. Thus, there is a need for more studies supporting the previous studies. Alpha-glucosidase was 1.5 times overexpressed in non-insulin-dependent diabetes mellitus patients, which increased postprandial glucose levels, according to reports [9]. The body absorbs monosaccharides and polysaccharides produced by α-amylase and α-glucosidase at varying rates, with monosaccharide components being absorbed very rapidly. It follows that inhibiting α-amylase and α-glucosidase action can delay the release of glucose from complex carbs, modifying the start of postprandial hyperglycemia, making it an excellent target for the treatment or management of type 2 diabetes mellitus [10]. Acarbose, a pseudo-carbohydrate derived from actinomycetes, is the α-glucosidase inhibitor that is most frequently administered [11]. In the present years, a quantity of bioactive compounds and their analogues from cyanobacteria with promising antibacterial, antiviral, anticancer, antifungal, antioxidant, anti-inflammatory, antidiabetic and anthel- mintic potentials have been explored [12].
Novel development for medications in all major disease categories can be found in natural resources, which constitute a significant source. Recently, the main research focus has been on generating novel medications using cyanobacteria [13]. They are present throughout the world, from the Antarctic to hot springs, deserts, soils, salt lakes, and brine water [14]. They serve as models for photosynthetic prokaryotes for the study of numerous metabolites produced by these organisms under such conditions due to their capacity to develop in a variety of ecological conditions, including light, temperature, salinity, alkalinity, and pollution [15-17]. According to studies [18, 19], the biologically active chemicals extracted from cyanobacteria exhibit antibacterial, antiviral, antifungal, enzyme inhibitory, immune-stimulant, cytotoxic, and anti-plasmodium effects. Additionally, cyanobacteria are a good source of additional vital compounds such as vitamins, amino acids, and fatty acids [20]. Biochemically active chemicals have been produced by cyanobacteria like Microcystis, Anabaena, Nostoc, Spirulina sp. and, Oscillatoriaetc [21]. The secondary metabolites produced by cyanobacteria are novel bioactive constituents used in the production of pharmaceuticals and vital chemicals for agriculture and various fields. It has been discovered that cyanobacterial lipopeptides are intriguing, biochemically active constituents. An enormous 85% of them are bioactive, including cytotoxic (41%), anticancerous (13%), antibiotics (12%), enzyme inhibitors (8%), antiviral (4%) and antifungal (4%) among their prominent properties [22]. The outstanding activity (18%) includes UV-absorbing properties that can be usable as sunscreens, herbicides, antimalarial, antimycotic, antimitotic, anti microalgal and activity that promotes cell differentiation [22].
Through the use of an in-silico approach, it is possible to identify lead compounds with the best binding energies and affinities for further study. These methods offer insightful information on the molecular interactions between bioactive constituents and target enzymes [23]. This study aims to investigate the prominent role of bioactive compounds derived from cyanobacteria in the treatment of type 2 diabetes mellitus, concentrating on how they interact with the enzymes α-amylase and α-glucosidase. The use of In-silico methods is to examine the affinities and binding energies of the aforementioned cyanobacterial bioactive compounds to the active sites of these enzymes. The research will clarify the inhibitory capability of compounds from cyanobacteria and their ability to control glucose release and carbohydrate digestion, helping to manage type 2 diabetes mellitus [24].
2. MATERIALS AND METHODS
2.1. Software and Tools
The recent study was executed by using bioinformatics tools and software like Auto Dock 4.2, Cygwin, and Discovery Studio 3.0.Operating systems that support 64-bit Windows 10 and 32-bit Windows 7 include biological databases like PubChem (http://pubchem.ncbi.nlm.nih. gov/), Protein Data Bank (www.rcsb.org/), Molsoft (https://www.molsoft.com/) and Pre ADMET (https://preadmet.webservice. bmdrc.org/).
2.2. Retrieval and Preparation of Targeted Protein Structures
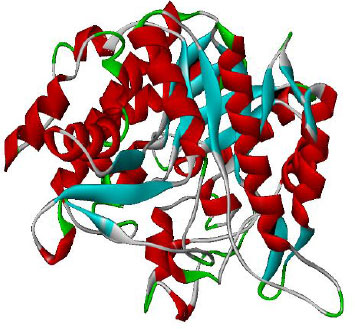
The main source for proteins having experimentally verified structures is the RCSB PDB (Research Collaboratory for Structural Bioinformatics, Protein Data Bank). It contains a large number of 3-dimensional structures that were discovered by techniques like NMR, X-ray crystallography, etc. The PDB was used to retrieve the targeted crystal structure of the proteins PDB ID: 1Kb3 (α- amylase) and 1qOX (α-glucosidase) shown in Figs. (1 and 2) from RCSB PDB (https://www.rcsb.org/.). The required hydrogen atoms were added after all water molecules, and crystallographic substructures from the specified protein were taken out to nullify the overall charges for molecular docking purposes. Furthermore, Discovery Studio Visualizer 3.0 was used to visualize the final structure, which shows the 3-D structure of selected proteins.

2.3. Ligand Library Preparation
All filtered derivatives that were retrieved in 3D sdf format from the PubChem database are included in the compound library. The library includes several hundred compounds that have been screened using Lipinski's rule of five and ADME filters. The drug molecules or chemical compounds' absorption, distribution, metabolism, excretion, and toxicity inside the animal body are represented by the ADME and toxicity profile. The blood-brain barrier, CaCo2, and AMES Toxicity are among the characteristics that make up the ADME profile. The network of blood arteries and tissues known as the blood-brain barrier protects the brain from hazardous constituents. The drug's enormous concentration will make it extremely difficult and even impossible for the human body to survive if it crosses the BBB. Human intestinal absorption (HIA) is a characteristic that shows that medications taken orally are absorbed into the bloodstream through the gastrointestinal tract. For a medication to be therapeutically effective inside the human body, it must have an HIA characteristic. The Ames test is a frequently used method for assessing a substance's genotoxicity by measuring its potential harm to human health through the identification of mutagenic characteristics. This test is defined by its exceptional repeatability and limitless sensitivity. We found 25 compounds with good ADME and toxicity characteristics after filtering.
2.4. Screening and Retrieval of Cyanobacterial and Control Compounds with Chemical Structures
A public chemical database called PubChem (https://pubchem.ncbi.nlm.nih.gov) was developed by the National Library of Medicine (NLM), a division of the National Institutes of Health (NIH) in the United States [25 Hundreds of bioactive compounds from different cyanobacterial strains selected by the literature survey were downloaded from PubChem database for the molecular docking studies and their chemical structures. Selected bioactive compounds filtered based on Lipinski’s rule of five (R05) and ADMET profile. Twenty-five cyanobacterial bioactive compounds selected out of 100, namely are - Acarbose (CID: 6918537), Metformin (CID: 4091), Ambiguine B Isonitrile (CID: 16109784), Ambiguine H Isonitrile (CID: 16069590), Ambigol A (CID: 475341), Ambigol B (CID: 475342), Anatoxin-a (CID: 3034748), Aplysiatoxin (CID: 46173823), Carazostatin (CID: 130857), Carbamydocyclophane E (CID: 16216032), Carbamy- docyclophane J (CID: 16216032), Curacin A (CID: 5281967), Curacin D (CID: 5281967), Debromoplysiatoxin (CID: 5352033), Nostocarboline (CID: 5326150), Ethyl Tumonoate A (CID: 53493313), Nostocine A (CID: 10749358), Nostocyclyne A (CID: 10088786), Radiosumin (CID: 102506248), Rutin (CID: 5280805), Tanikolide (CID: 5276592), Venturamide B (CID: 16115401) Scytoscalarol (CID: 44605340) and also other compounds with chemical structures retrieved from ‘PubChem’ database are shown in Table 1.
| S.No. | Compounds Name | PubChem No. | Source of Compounds | Compounds Structure |
|---|---|---|---|---|
| 1 | METFORMIN (CONTROL) | 4091 | Galega officinalis |
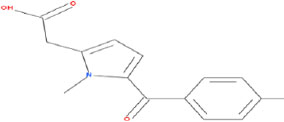
|
| 2 |
ACARBOSE
(CONTROL) |
41774 | Synthetic drug |
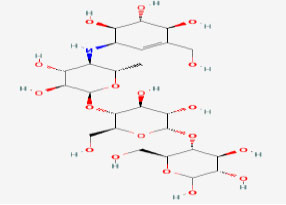
|
| 3 | Ambiguine I isonitrile | 16109784 | Fischerella sp. |
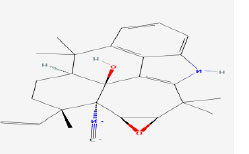
|
| 4 | Ambiguine H isonitrile | 16069590 | Fischerella sp. |
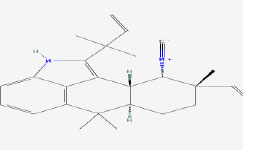
|
| 5 | Ambigol A | 475341 | Fischerella ambigua |
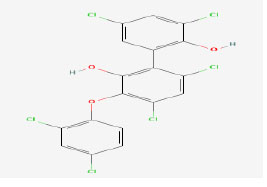
|
| 6 | Ambigol B | 475342 | Fischerella ambigua |
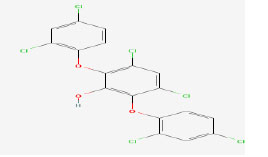
|
| 7 | Anatoxin-a | 431734 | Anabaena circinalis |
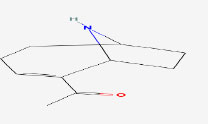
|
| 8 | Aplysiatoxin | 21672114 | Geitlerinema |
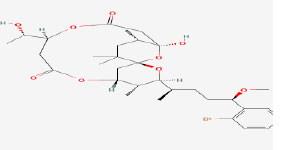
|
| 9 | Carazostatin | 130857 | Hyella caespitose |
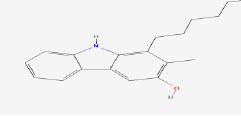
|
| 10 | Carbamidocyclophane E | 16216032 | Nostoc sp. |
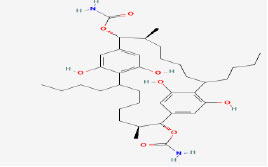
|
| 11 | Carbamidocyclophane J | 122206459 | Nostoc sp. |
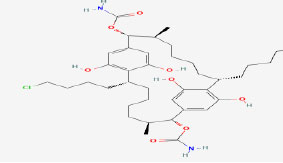
|
| 12 | Curacin D | 10546404 | Lyngbya sp. |
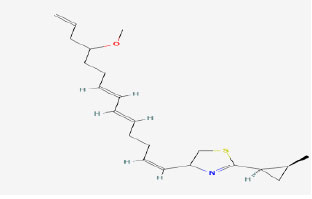
|
| 13 | Curacin A | 5281967 | Lyngbya sp. |
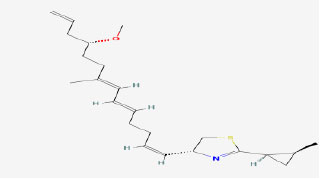
|
| 14 | Cylindrospermopsin | 42628600 | Cylindrospermopsis sp. |
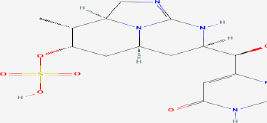
|
| 15 | Debromoaplysiatoxin | 5352033 | Lyngbya majuscule |
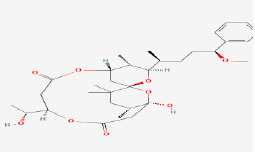
|
| 16 | Ethyl Tumonoate A | 53493313 | Oscillatoria margaritifera |
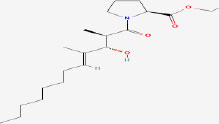
|
| 17 | Nostocarboline | 5326150 | Nostoc sp. |
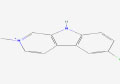
|
| 18 | Nostocine A | 10749358 | Nostoc spongiae forme |
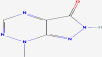
|
| 19 | Nostocycline A | 10088786 | Nostoc sp. |
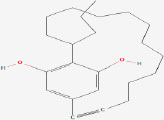
|
| 20 | Radiosumin | 10812638 | Plectonema radiosum |
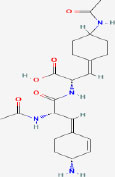
|
| 21 | Rutin | 5280805 | Gracilaria dendroides |
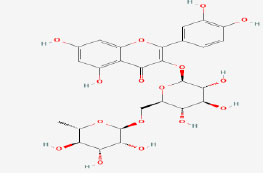
|
| 22 | Scytoscalalrol | 44605340 | Scytonema pseudohofmanni |
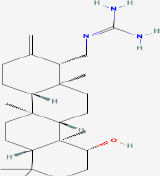
|
| 23 | Tanikolide | 5276592 | Lyngbya majuscula |

|
| 24 | Tjipanazole J | 15705741 | Tolypothrix tjipanasensis |
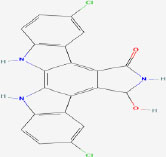
|
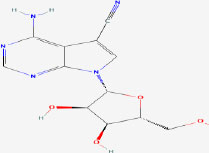
| 25 | Toyocamycin | 11824 | Tolypothrix tenuis |
|
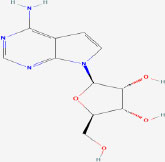
| 26 | Tubercidin | 6245 | Plectonema radiosum |
|
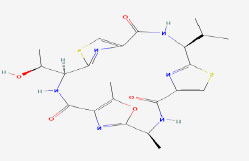
| 27 | Venturamide B | 16115401 | Oscillatoria sp. |
|
2.5. Prediction of ADME and Toxicity Profile
All 100 of the resulting compounds were subsequently approved to go through ADME and toxicity filtration after the Lipinski filter had been applied. All 100 cyanobacterial compounds' ADMET properties were predicted using the preADMET tool, and only candidates that favoured the values of the BBB, CaCo2, HIA, and AMES toxicity and Mouse/Rat carcinogenicity parameters were chosen. Out of 100 compounds, 25 were found to have useful ADMET profiles after the ADMET screening. The same preADMET web server was used to evaluate the toxicity profile of the 25 compounds that were chosen.
2.6. Molecular Docking using AutoDock 4.2 and Cygwin
The bioinformatics tools AutoDock 4.2 and Cygwin were used to evaluate and identify the lowest free binding energy of the selected cyanobacterial compounds along with 2 reference compounds against the targeted protein α- amylase (1Kb3) and α-glucosidase (1qOX) which is associated with type 2 diabetes mellitus.
3. RESULTS AND DISCUSSION
3.1. Drug-likeness Properties of Cyanobacterial Compounds
The main purpose of Lipinski's rule of five, commonly referred to as the thumb rule, is to assess the drug-likeness of a chemical compound in order to establish whether or not a compound with a particular pharmacological activity is orally active in humans. For a molecule to function as a medication and demonstrate good permeability through cell membranes, the log P value must be lower than 5. Other criteria for this rule state that the number of hydrogen bond acceptors must be ≤ 10 (The sum of Os and Ns), the number of hydrogen bond donors must be ≤ 5 (The sum of OHs and NHs), and molecular weight should be <500Da. If the number of violations are equal to 1 or no violation, it states that the compound can simply bind to the target receptor [26]. Drugs are excluded from further screening if more than two parameters are out of the standard range [27]. Out of 100 screened compounds from cyanobacteria, only 25 compounds were filtered through Lipinski’s Rule of five and were taken further for ADMET profiling. The list of filtered compounds is given in Table 2.
3.2. ADMET Evaluation of Filtered Compounds
According to Table 3, 25 filtered compounds from Lipinski's rule of five were used in the ADMET investigation. Studies show that poor pharmacokinetics and toxicity are the main reasons for expensive late-stage drug development failures, and it is now generally recognized that these factors should be taken into account as early as feasible in the drug discovery process. The number of compounds for which preliminary data on absorption, distribution, metabolism, excretion (ADME), and toxicity (T) are required has greatly expanded as a result of combinatorial chemistry and high-throughput screening [27]. PreADMET, a tool that is accessible online (https://preadmet.webservice.bmdrc.org), evaluates ADME attributes.
| S. No. | Name of the Compounds | Drug Likeness Properties | Drug- Likeness Score | |||
|---|---|---|---|---|---|---|
| LogP | Molecular Weight | Hydrogen Bond Donor | Hydrogen Bond Acceptor | |||
| A | METFORMIN (CONTROL) | -1.00 | 129.10 | 5 | 2 | -0.82 |
| B |
ACARBOSE
(CONTROL) |
-4.58 | 645.25 | 14 | 19 | 0.40 |
| 1 | Ambiguine I Isonitrile | 6.19 | 402.50 | 2 | 2 | -1.1 |
| 2 | Ambiguine H Isonitrile | 7.58 | 372.26 | 1 | 1 | -0.78 |
| 3 | Ambigol A | 8.36 | 485.00 | 2 | 3 | -0.14 |
| 4 | Ambigol B | 8.16 | 485.00 | 1 | 3 | -0.48 |
| 5 | Anatoxin-a | 1.53 | 165.23 | 1 | 1 | -1.42 |
| 6 | Aplysiatoxin | 4.93 | 670.24 | 3 | 10 | 0.32 |
| 7 | Carazostatin | 6.83 | 295.40 | 2 | 1 | -0.41 |
| 8 | Carbamydocyclophane E | 8.71 | 670.90 | 8 | 8 | 0.82 |
| 9 | Carbamydocyclophane J | 8.71 | 670.90 | 8 | 8 | 0.82 |
| 10 | Curacin A | 6.51 | 373.60 | 0 | 3 | 0.35 |
| 11 | Curacin D | 6.51 | 373.60 | 0 | 3 | 0.35 |
| 12 | Debromoplysiatoxin | 4.33 | 592.70 | 3 | 10 | 0.47 |
| 13 | Nostocarboline | 3.36 | 217.67 | 1 | 0 | -0.94 |
| 14 | Ethyl Tumonoate A | 5.08 | 367.50 | 1 | 4 | -0.60 |
| 15 | Nostocine A | -0.92 | 151.13 | 1 | 4 | -1.16 |
| 16 | Nostocyclyne A | 8.36 | 342.50 | 2 | 2 | -0.08 |
| 17 | Radiosumin | -1.95 | 432.50 | 6 | 6 | -0.21 |
| 18 | Rutin | -1.55 | 610.50 | 10 | 16 | 0.91 |
| 19 | Scytoscalalrol | 4.83 | 415.7 | 5 | 2 | -0.47 |
| 20 | Tanikolide | 5.34 | 284.40 | 1 | 3 | -0.95 |
| 21 | Tjipanazole J | 5.94 | 290.7 | 2 | 0 | -1.31 |
| 22 | Tolyporphin | 4.08 | 742.8 | 4 | 12 | 0.02 |
| 23 | Toyocamycin | -1.02 | 291.26 | 5 | 7 | 0.53 |
| 24 | Tubercidin | -0.92 | 266.25 | 5 | 6 | 0.54 |
| 25 | Venturamide B | 1.75 | 518.60 | 4 | 10 | -0.58 |
| S. No. | Name of The Compound | BBB | HIA | Caco2 | AMES Toxicity |
Carcinogenicity Mouse/rat |
|---|---|---|---|---|---|---|
| 1 | Ambiguine I Isonitrile | 7.45316** | 92.849612 | 23.9099 | Mutagen | Positive/ Positive |
| 2 | Ambiguine H isonitriles | 16.5872 | 97.046579 | 48.6152 | Mutagen | Positive/ Negative |
| 3 | Ambigol A | 13.4989 | 95.546983 | 46.5271 | Mutagen | Negative/Positive |
| 4 | Ambigol B | 14.9377 | 96.842893 | 56.983 | Mutagen | Negative/Positive |
| 5 | Anatoxin-a | 0.612474 | 95.398255 | 30.5172 | Mutagen | Positive/ Negative |
| 6 | Aplysiatoxin | 0.62309 | 93.927517 | 21.7641 | Non-Mutagen | Negative/Negative |
| 7 | Carazostatin | 15.6199 | 93.267025 | 45.6263 | Mutagen | Negative/Negative |
| 8 | Carbamidocyclophane E | 6.6225 | 78.754511 | 20.4097 | Mutagen | Negative/Negative |
| 9 | Carbamidocyclophane J | 6.54959 | 85.988582 | 21.0478 | Mutagen | Negative/Negative |
| 10 | Curacin D | 3.20083 | 97.506753 | 31.6657 | Mutagen | Positive/ Negative |
| 11 | Curacin A | 5.7525 | 97.546286 | 35.2846 | Non-Mutagen | Positive/ Negative |
| 12 | Cylindrospermopsin | 0.0412124 | 69.533771 | 21.0569 | Mutagen | Negative/Positive |
| 13 | Debromoaplysiatoxin | 1.92282 | 96.111782 | 33.1237 | Non-Mutagen | Negative/Negative |
| 14 | Ethyl Tumonoate A | 0.239313 | 90.065359 | 24.9041 | Mutagen | Positive/Negative |
| 15 | Nostocarboline | 5.18456 | 97.910575 | 16.7822 | Mutagen | Positive/Negative |
| 16 | Nostocine A | 0.114421 | 78.765338 | 11.6462 | Mutagen | Positive/Positive |
| 17 | Nostocycline A | 18.0699 | 93.610401 | 47.7531 | Mutagen | Negative/Negative |
| 18 | Radiosumin | 0.0598323 | 63.980144 | 20.8546 | Mutagen | Positive/Positive |
| 19 | Rutin | 0.0285642 | 2.861176 | 7.91267 | Non-Mutagen | Negative/Negative |
| 20 | Scytoscalarol | 0.75716 | 92.010946 | 21.0215 | Mutagen | Negative/ Positive |
| 21 | Tanikolide | 7.06301 | 94.877159 | 46.7781 | Non-Mutagen | Negative/Negative |
| 22 | Tolyporphin J | 0.067607 | 82.169793 | 0.808384 | Non-Mutagen | Positive/Positive |
| 23 | Toyocamycin | 0.112285 | 46.547342 | 5.36832 | Mutagen | Negative/ Positive |
| 24 | Tubercidin | 0.360937 | 65.797783 | 5.22828 | Mutagen | Negative/Negative |
| 25 | Venturamide B | 0.0561416 | 56.779147 | 9.19956 | Mutagen | Negative/ Positive |
A total of 25 ligands from Cyanobacteria were selected out of 100 for the molecular docking investigation. After molecular docking investigation of selected ligands out of 25 top 3 compounds, namely, Ethyl Tumonoate A, Debromoaplysiatoxin and Scytoscalarol, showed highly significant interaction with α-amylase listed in Table 4 and also out of 25 ligands top 3 compounds namely, Ambiguine I isonitrile, Cylindrospermopsin and Debromoaplysiatoxin showed excellent binding interaction with α-glucosidase listed in Tables 5 and 6.
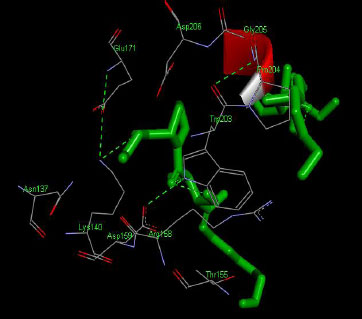
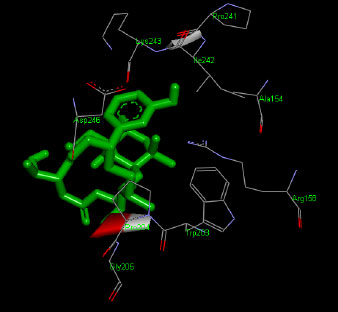
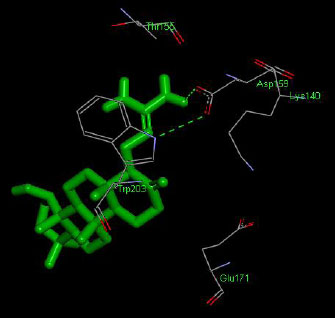
The molecular docking interaction results reveal the free binding energy of α-amylase was found with Metformin (reference compound) concluded 10 amino acid residues, namely, Ile 235, Glu 233, Ala 193, Asp 197, Asp 96, Tyr 62, Trp 58, His 299, Asp 300, His 305 (Fig. 3); with Ehtyl Tumunoate A concluded 10 amino acid residues- Glu 171, Asp 206, Gly 205, Pro 204, Trp 203, Asn 137, Lys 140, Asp 159, Arg 158, Thr 155 (Fig. 4); with Debromoaplysiatoxin concluded 07 amino acid residue- Pro 241, Lys 243, Ala 154, Asp 246, Gly 205, Trp 203, Arg 158 (Fig. 5); with Scytoscalarol concluded 05 amino acid residues- Thr 155, Asp 159, Lys 140, Trp 203, Glu 171 (Fig. 6). The values of free binding energy for Metformin with α-amylase, Ehtyl Tumunoate A with α-amylase, and Scytoscalarol with α-amylase docking scores originated were –7.66, –9.28, –7.97, -7.14 kcal/mol respectively, although the conforming values of inhibition constant (Ki) were expected to be 2.43, 0.5765, 1.43, 5.80 µM separately listed in Table 5.
Figs. (3-6): Docked structure of selected top three leads and Metformin, a control drug, with the active site residues of α-amylase.
| S. No. | Name of Compounds |
Free Binding Energy Kcal/mol |
Inhibition Constant Ki (μM) |
|---|---|---|---|
| 0 | METFORMIN (Control) | -7.66 | 2.43 |
| 1 | Ethyl Tumonoate A | -9.28 | 0.15765 |
| 2 | Debromoaplysiatoxin | -7.97 | 1.43 |
| 3 | Scytoscalarol | -7.14 | 5.80 |
| 4 | Ambigol A | -4.93 | 241.67 |
| 5 | Ambigol B | -5.21 | 152.01 |
| 6 | Ambiguine H isonitriles | -4.95 | 235.76 |
| 7 | Tanikolide | -3.28 | 3910 |
| 8 | Tjipanazole J | -5.63 | 75.05 |
| 9 | Tolyporphin J | -6.07 | 35.63 |
| 10 | Toyocamycin | -5.27 | 116.27 |
| 11 | Tubercidin | -4.62 | 411.82 |
| 12 | Venturamide B | -5.02 | 209.47 |
| 13 | Nostocarboline | -4.45 | 550.91 |
| 14 | Nostocine A | -4.56 | 493.97 |
| 15 | Nostocycline A | -6.21 | 28.07 |
| 16 | Radiosumin | -5.2 | 153.42 |
| 17 | Rutin | -4.39 | 606.76 |
| 18 | Carbamidocyclophane E | -4.13 | 940.81 |
| 19 | Carbamidocyclophane J | -3.34 | 3.58 |
| 20 | Curacin D | -6.31 | 23.89 |
| 21 | Curacin A | -3.23 | 4270 |
| 22 | Ambiguine B isonitrile | -5.17 | 162.87 |
| 23 | Anatoxin-a | -5.00 | 217.36 |
| 24 | Aplysiatoxin | -5.49 | 95.04 |
| 25 | Carazostatin | -5.20 | 155.48 |
| S. No. | Name of Compounds |
Free Binding Energy Kcal/mol |
Interacting Residues in the Receptor |
|---|---|---|---|
| 0 | METFORMIN (CONTROL) | -7.66 | Ile 235, Glu 233, Ala 193, Asp 197, Asp 96, Tyr 62, Trp 58, His 299, Asp 300, His 305 |
| 1 | Ethyl Tumonoate A | -9.28 | Glu 171, Asp 206, Gly 205, Pro 204, Trp 203, Asn 137, Lys 140, Asp 159, Arg 158, Thr 155 |
| 2 | Debromoaplysiatoxin | -7.97 | Pro 241, Lys 243, Ala 154, Asp 246, Gly 205, Trp 203, Arg 158 |
| 3 | Scytoscalarol | -7.14 | Thr 155, Asp 159, Lys 140, Trp 203, Glu 171 |
| S. No. | Name of Compounds |
Free Binding Energy Kcal/mol |
Inhibition Constant Ki (μM) |
|---|---|---|---|
| 0 | ACARBOSE (control) | -8.86 | 0.31974 |
| 1 | Ambiguine I isonitriles | -9.46 | 0.11641 |
| 2 | Scytoscalarol | -7.5 | 3.2 |
| 3 | Cylindrospermopsin | -7.17 | 5.51 |
| 4 | Rutin | -2.01 | 33410 |
| 5 | Ambiguine H isonitriles | -4.97 | 226.79 |
| 6 | Tanikolide | -5.05 | 198.48 |
| 7 | Tjipanazole J | -6.18 | 29.62 |
| 8 | Toyocamycin | -6.59 | 14.68 |
| 9 | Venturamide B | -5.79 | 57.3 |
| 10 | Debromoaplysiatoxin | -4.94 | 241.16 |
| 11 | Ethyl Tumonoate A | -3.47 | 2870 |
| 12 | Nostocarboline | -4.51 | 497.92 |
| 13 | Nostocine A | -4.57 | 445.18 |
| 14 | Nostocycline A | -4.97 | 227.14 |
| 15 | Radiosumin | -4.88 | 267.02 |
| 16 | Carbamidocyclophane J | -2.42 | 16750 |
| 17 | carbamidocyclophane c | -2.42 | 16780 |
| 18 | Ambigol A | -5.82 | 54.22 |
| 19 | Ambigol B | -6.55 | 15.72 |
| 20 | Anatoxin-a | -6.10 | 33.65 |
| 21 | Aplysiatoxin | -4.52 | 490.20 |
| 22 | Carazostatin | -6.07 | 35.76 |
| 23 | Carbamidocyclophane E | -2.94 | 7.04 |
| 24 | Curacin D | -4.70 | 1.03 |
| 25 | Curacin A | -4.24 | 776.45 |
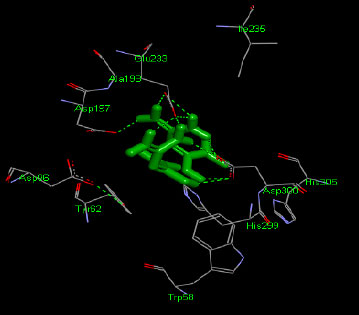
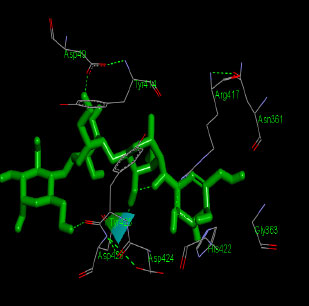
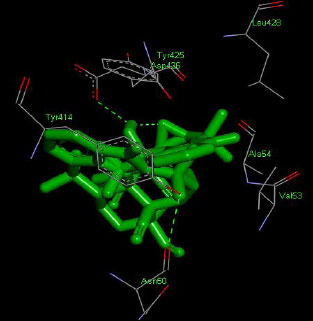
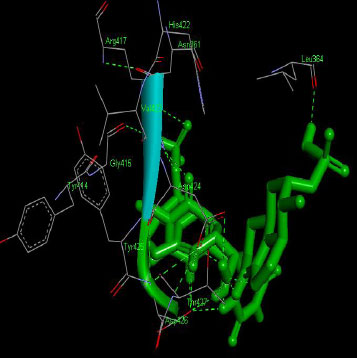
Similarly, the molecular docking interaction revealed that the free binding energy of α-glucosidase was found with Acarbose (reference compound) showed 09 amino acid residues-Asp 49, Tyr 414, Arg 417, Asn 361, Tyr 425, Asp 426, Asp 424, His 422, Gly 363 (Fig. 7); with Ambiguine I Isonitrile showed 07 amino acid residues-Leu 428, Tyr 425, Asp 426, Tyr 414, Ala 54, Val 53, Asn 50 (Fig. 8); with Scytoscalarol showed 09 amino acid residues - Asp 49, Gly 415, Tyr 414, Asp 426, Val 53, Tyr 425, Asp 424, Arg 417, Leu 428 (Fig. 9); with Cylindrospermopsin showed 11 amino acid residues - Arg 417, His 422, Asn 361, Leu 364, Val 423, Gly 415, Tyr 414, Asp 424, Tyr 425, Thr 427, Asp 426 (Fig. 10). The values of free binding energy for Acarbose with α-glucosidase; Ambiguine I Isonitrile with α-glucosidase; Scytoscalarol with α-glucosidase; Cylindrospermopsin with α-glucosidase are “–8.86, –9.46, –7.5, -7.17 kcal/mol respectively, although the conforming values of inhibition constant (Ki) were expected to be 0.31974; 0.1164; 3.20; 5.51 µM which are separately listed in Table 7.
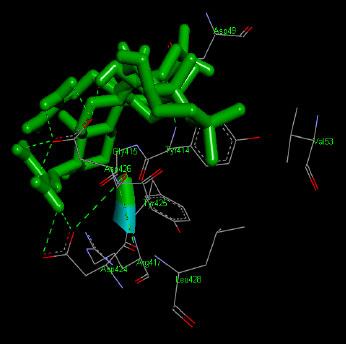
Figs. (7-10): Docked structure of selected top three leads and Acarbose, a control drug, with the active site residues of α-glucosidase.
| S. No. | Name of Compounds |
Free Binding Energy Kcal/mol |
Interacting Residues in the Receptor |
|---|---|---|---|
| 0 | ACARBOSE (Reference compound) | -8.86 | Asp 49, Tyr 414, Arg 417, Asn 361, Tyr 425, Asp 426, Asp 424, His 422, Gly 363 |
| 1 | Ambiguine I isonitriles | -9.46 | Leu 428, Tyr 425, Asp 426, Tyr 414, Ala 54, Val 53, Asn 50 |
| 2 | Scytoscalarol | -7.5 | Asp 49, Gly 415, Tyr 414, Asp 426, Val 53, Tyr 425, Asp 424, Arg 417, Leu 428 |
| 3 | Cylindrospermopsin | -7.17 | Arg 417, His 422, Asn 361, Leu 364, Val 423, Gly 415, Tyr 414, Asp 424, Tyr 425, Thr 427, Asp 426 |
The resultant compounds were also found to be suitable with noble druggable character, as per Lipinski’s rule of five and ADEMT profile properties. Therefore, this potent compound could be a possible drug applicant for treating type 2 diabetes mellitus [28]. The resulting cyanobacterial compounds are subjected to in-vitro analysis where phytochemical evaluation, anti-oxidant activity, anti-diabetic activity and anti-inflammatory activity. Current progress in the understanding of the biology of T2DM has resulted in an increasing number of therapies and remedies that are permitted or in clinical development for this disorder [29]. Some compounds have exhibited very interesting results and successfully reached Phase II and Phase III clinical trials [30].
CONCLUSION
Diabetes mellitus is a global metabolic disorder that is alarmingly on the rise. The prolonged use of currently available synthetic medications offers only a temporary solution and comes with undesirable side effects. The resurgence of the natural medicine field holds promise, and there is a strong call for solutions to diabetes from natural sources. Recently, there have been reports highlighting cyanobacteria as a valuable source of bio- active compounds, including Ambiguine I Isonitrile, Cylindrospermopsin, Scytoscalalrol, Ambigol A, Ambigol B, Anatoxin-a, Aplysiatoxin, Carazostatin, Carbamydo- cyclophane E, Carbamydocyclophane J, Curacin A, Curacin D, Debromoplysiatoxin, Nostocarboline, Ethyl Tumonoate A, Nostocine A, Nostocyclyne A, Radiosumin, Rutin, Scytoscalarol, Tanikolide, and Venturamide A.
Furthermore, these bioactive compounds from cyanobacteria hold immense potential in research and offer a limitless outlook for the development of new drug candidates. In this study, molecular docking analysis has unveiled that certain cyanobacterial bioactive compounds, such as Ambiguine I Isonitrile, Cylindrospermopsin, Scytoscalalrol, and Ethyl Tumonoate A, exhibit strong potential as candidates for the development of novel drugs targeting α-amylase and α-glucosidase enzymes in the treatment of type 2 diabetes mellitus. Further experimental in-vitro and in-vivo studies are needed to confirm the therapeutic effectiveness of these compounds in the development of innovative antidiabetic medications.
LIST OF ABBREVIATIONS
| T2DM | = Type 2 Diabetes Mellitus |
| DM | = Diabetes Mellitus |
| WHO | = World Health Organization |
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
Not applicable.
HUMAN AND ANIMAL RIGHTS
Not applicable.
CONSENT FOR PUBLICATION
Not applicable.
AVAILABILITY OF DATA AND MATERIALS
The data and supportive information are available within the article.
FUNDING
None.
CONFLICT OF INTEREST
Salman Akhtar is the Associate Editorial Board Member of The Open Bioinformatics Journal.
ACKNOWLEDGEMENTS
The authors would like to thank Integral University, Lucknow, for providing the facilities to carry out this research work, with Manuscript Communication Number: IU/R&D/2023 MCN0002167.


