Virtual Screening and Molecular Simulation of Drug Candidates Targeting the Human SLC4A4 Protein for Colorectal Cancer Therapy
Abstract
Introduction
Colorectal cancer is a highly complex disease that continues to rise in prevalence, posing significant difficulties in its management and treatment outcomes. Reduced expression of the SLC4A4 gene has been identified as a critical factor in driving tumor development and poor clinical prognosis in CRC. This reduction disrupts several key cellular mechanisms, including cellular proliferation, programmed cell death, and metastasis, largely by altering pH regulation within the cells. Given its influence on tumor behavior and patient survival, SLC4A4 expression levels could serve as a reliable prognostic marker in colorectal cancer.
Methods
The CryoEM structure of SLC4A4, retrieved from the Protein Data Bank (PDB), was refined using SwissModel and subjected to virtual drug screening via the DrugRep web server. This screening employed a database comprising FDA-approved drugs.
Results
Nilotinib emerged as the most potent inhibitor following further validation through redocking using CBDOCK2, ranking it highest among the top three results from DrugRep. The docking analysis yielded a score of −11.2 kcal/mol for Nilotinib. A 50-nanosecond molecular dynamics simulation further validated these findings, revealing that Nilotinib formed robust interactions with the SLC4A4 protein. The simulation yielded consistent results, with a root mean square deviation of around 0.30 nm, a root mean square fluctuation of near 0.5 nm, a compact radius of gyration between 3.8 and 4.0 nm, and stable solvent-accessible surface area profiles. These findings confirmed that the drug-protein complex maintained structural stability throughout the simulation.
Discussion
The computational findings suggest that Nilotinib binds effectively and stably to SLC4A4, indicating its potential to modulate the function of this protein in CRC. Its established safety profile as an FDA-approved drug supports the feasibility of drug repurposing. These results also reinforce the utility of integrating structure-based drug screening with molecular dynamics simulations to identify novel therapeutic agents for cancer.
Conclusion
Nilotinib holds significant potential as a therapeutic agent for colorectal cancer and warrants further experimental investigation to validate its effectiveness.
1. INTRODUCTION
Colorectal cancer (CRC) is among the most common and lethal malignancies worldwide, leading to an increasing percentage in cancer-related disease incidence and patient deaths [1, 2].In this new genomic era, the understanding of CRC has shifted pretty dramatically. There’s a layered genome at play that drives tumor growth and biology [3]. Even with solid advances in diagnosing and treating colon cancer, the 5-year survival rate for advanced stages stays alarmingly low, less than 30%, mostly because of recurrence and metastasis after surgery. This has prompted a deeper dive into the mechanisms underlying colon cancer progression and a need to identify new biomarkers to enhance personalized treatment strategies and prognostic evaluations for patients [4]. There’s growing evidence that abnormal metabolism is a pervasive feature of tumor development, and it significantly influences how tumors progress and resist therapy [5-7]. The final product of glycolysis, i.e., Pyruvate, plays a crucial role in both cellular anabolism and catabolism. Turns out, metabolic misregulation in tumor cells isn't just about turning certain signaling pathways on or off; it's also about how driver genes get regulated, think those involved in partial epithelial-mesenchymal transition (p-EMT) [8-11]. Tumor metabolites are involved in core functions such as lipid uptake, synthesis, and hydrolysis, and they influence the biological behavior of malignant tumors. Hence, examining genes involved in pyruvate metabolism might offer valuable insights for developing targeted tumor therapies [12, 13].
Analyzing gene expression profiles across different diseases is essential for deciphering underlying molecular mechanisms and identifying potential therapeutic targets [14]. For example, the Expression Atlas provides analyses of gene/protein expression in tissues, types, and diseases [15].
A gene of interest in CRC is the solute carrier family 4, member 4 (SLC4A4) gene [16]. Recent studies have shown that downregulation of SLC4A4 is correlated with Epithelial-Mesenchymal Transition (EMT) and metastasis in CRC [17]. SLC4A4 can be associated with immune cell infiltration (CD8+T cells, dendritic cells, and NK cells), further supporting the tumor microenvironment [18]. Cappellesso et al. (2022) reported that SLC4A4 can confer resistance to immunotherapy, and it is a prognostic biomarker and therapeutic target [19]. Together, these findings show that SLC4A4 does more than regulate pH; it may also help suppress tumor growth and represent a potential therapeutic target.
SLC4A4 encodes a sodium bicarbonate cotransporter that is important for cellular pH homeostasis [20, 21]. Abnormal expression of SLC4A4 has been reported in several malignancies, including CRC [22]. Tumor microenvironment pH regulation is critical for the survival and proliferation of cancer cells, and SLC4A4 may represent a novel candidate for intervention [19, 23]. By targeting SLC4A4 function and intracellular pH, the proliferation and metastasis of CRC cells may be retarded, revealing novel therapeutic approaches [24]. Ion transporters, including those of the SLC family, have previously been shown to alter cancer cell growth and invasion [25, 26]. SLC4A4 acts as a tumor suppressor gene and is significantly downregulated and associated with microsatellite instability [18].
In the pursuit of identifying effective treatments for CRC, drug repurposing has emerged as a viable strategy [27]. This is a process of identifying existing FDA-approved drugs that could be repurposed to target the new molecular pathways identified in CRC [28]. Virtual screening of drug databases targeting the SLC4A4 refined structure may facilitate the identification of potential inhibitors [29].
With the rich genetic and functional information available through Expression Atlas, novel therapeutic targets can be identified to hasten the development of CRC treatments. This integrated strategy will not only advance our knowledge of CRC biology but will also guide personalized medicine, in which therapeutic interventions are tailored to the genetic composition of individual tumours.
Taken together, the Expression Atlas data, the importance of SLC4A4 in CRC, and the drug repurposing approach constitute an opportunity to develop a precision medicine therapeutic. With the current study of the molecular machinery of CRC and the repositioning of drugs, it is possible to identify a more promising, personalized drug candidate against an intractable disease.
2. MATERIAL AND METHODS
2.1. Data Acquisition
To investigate the expression pattern of the gene SLC4A4 in colorectal cancer (CRC), we accessed a public resource, Expression Atlas, established by EMBL-EBI (https://www.ebi.ac.uk/gxa/home) [15]. The gene-specific data were retrieved by searching for the gene symbol (SLC4A4) or the Ensembl gene ID (ENSG00000080493). From the gene overview page, we selected the “Differential expression” tab, which reports changes in gene expression across different experimental conditions and tissues. This tabular result contains major features such as experiment accession E-GEODseries, comparisons of disease versus control, log2FC, and p-value (adjusted p-value) [15]. Then, the findings were reanalyzed to identify fold changes in gene expression. It was performed by evaluating SLC4A4 expression in CRC tissues compared with normal tissues. The analysis was performed to confirm differential expression reliably enabling to analyze the role of SLC4A4 in CRC.
2.2. Protein Preparation
NBCe1, the human SLC4A4, a sodium-coupled acid-base transporter, was prioritized in this study from a pool of several colorectal cancer-associated gene expression data obtained from the Expression Atlas database. Homology modeling was performed to refine the CryoEM structure of the human SLC4A4 protein (PDB ID: 6CAA) [30] and to identify potential structural gaps/ missing amino acid residues for subsequent investigation using the SWISS-MODEL server [31, 32]. In this procedure, the SWISS-MODEL template library (SMTL 2024-03-27 version) was searched for evolutionarily related protein structures as templates. The best template is then selected based on a heuristic selection strategy.
2.3. Screening (Receptor-based)
For receptor-based screening, the 3D structure of the target protein (PDB ID: 6CAA) retrieved from SWISS-MODEL was used to find potential binding pockets on the protein surface. The DrugRep platform, an automated, parameter-free virtual screening server, was used to facilitate this process. DrugRep employs CurPocket, an in-house curvature-based cavity-detection method, to automatically predict potential pockets on the protein surface. This method is efficient for extracting docking boxes for molecular docking. Next, a high-throughput virtual screening was performed on a curated set of 2470 FDA-approved drugs from the DrugBank (version 5.1.7) collection. Docking procedure. The batch docking over the drug library was performed using DrugRep, which provides a procedure for batch docking using the widely used molecular docking tool AutoDock Vina (version 1.1.2) [33]. The platform sets the docking parameters, including the grid center coordinates and the dimensions of the search region, based on the predicted pockets.
The exhaustiveness parameter was set to 8, the default for DrugRep, which balances computational speed and accuracy. Docking results were ranked by Vina binding affinity scores in kcal/mol. Compounds with docking scores lower than –8.0 kcal/mol were considered promising hits for further analysis. This threshold was selected as a stringent cutoff to identify compounds with strong predicted binding affinities, and it is a common practice in virtual screening studies to filter a large library down to a manageable number of lead candidates [34]. The top hits were then passed to a more refined docking step with CBDock2 [35].
2.4. Curation and Preparation of FDA-Approved Ligands for Virtual Screening
The DrugRep cultivated approved drug library refers to a collection of FDA-approved drugs that have been modified for curation and made available in virtual screening campaigns or repurposing studies. This library includes current, up-to-date medications that are already FDA-approved for human use, along with detailed pharmacological profiles and safety data. As part of the high-throughput docking workflow, the online tool DrugRep was used with default parameters to prepare ligands from a curated library of FDA-approved drugs obtained from the DrugBank database (version 5.1.7).This intentional emphasis on a known library of FDA-approved drugs is one of the important advantages of our drug repurposing approach, as it obviates the need for the comprehensive safety and pharmacokinetic studies required for new chemical entities. The compounds were ranked by docking score, with higher scores indicating stronger predicted interactions with the protein's binding site. The three best-scoring candidates were taken forward for further studies to identify the best drug [34].
2.5. Molecular Docking Using CB-DOCK2
To enhance the docking analysis and verify the binding poses of the top 3 drugs (DB01396 [Digitoxigenin], DB00762 [Irinotecan], and DB04868 [nilotinib]), we used the CB-Dock2 web server. CB-Dock2, a two-tier blind docking algorithm with template-based and structure-based blind docking approaches. The server searches for homologous templates with similar protein and ligand structures to facilitate docking if the above first steps are successful. Yet, in this study, with the protein and ligands of interest, the CB-Dock2 server reported that no suitable templates were found in its database [35].
Therefore, all three ligand docking was performed via the ligand-protein blind docking protocol. This procedure, inherited from the original CB-Dock server, is based on the AutoDock Vina engine and guided by the surface curvature of a protein to find potential binding cavities. It further docks molecular structures within the identified pockets for detection to optimize ligand and ligand conformations and evaluate binding affinities. The server generated multiple binding poses for each ligand in some of the predicted cavities. Vina binding affinity scores (in kcal/mol) were used for the final ranking of the complexes, and the best-scored pose for each ligand was considered for subsequent analysis. This protocol successfully identified both the most likely binding sites and the types of interactions involved between ligands and an SLC4A4 protein [35].
2.6. Molecular Dynamics Simulation
The MD simulations were performed with GROMACS 2024.1- dev version (www.gromacs.org) in combination with the CHARMM36-jul2022.ff version of the force field under CHARMM36-jul2022.ff [36]. The initial protein-ligand complex conformation was achieved with CB-DOCK2. Protein and ligand structures were prepared using the pdb2gmx utility, and missing hydrogens were added during the process. Ligand topology and parameters files were created with the latest version of CGenFF through CHARMM-GUI [37]. Using the TIP3P water model, the simulation system was solvated in a cubic water box, with a 10 Å buffer between the protein and the box edges. Charge neutrality was achieved by adding sodium ions. Both systems were treated with periodic boundary conditions for a 50 ns run using the Leapfrog integrator, along with equilibration at NVT and NPT conditions. Electrostatics were calculated by ParticleMesh Ewald (PME). Invite a force-based switching function to capture non-bonded interactions over 10 and 12 Å. To remove the steric clashes and bad contacts, 50,000 steps of steepest descent were run for energy minimization. Structural analyses, such as RMSD and RMSF, were performed using gmxrms and gmxrmsf, respectively. Hydrogen bond analysis was performed with gmxhbond. Additionally, Rg and SASA were calculated using gmx gyrate and gmx sasa, respectively. Simulation data were visualized in PyMOL and plotted using Grace software [38, 39]. Ligand-receptor dynamics were simulated with 50 ns on the GridMarkets platform, which took ~22 hours.
2.7. Study Design
An exploratory quantitative and computational analysis is conducted that combines publicly available transcriptomic data with in silico drug screening tools. The main goal was to investigate the importance of the SLC4A4 gene in colorectal cancer (CRC) and to identify potential FDA-approved drugs targeting its protein product. The study was a structured multi-stage computational process:
- Gene Expression Analysis – SLC4A4 expression in CRC vs. normal tissue was assessed for differential expression using Expression Atlas public data sets, with log2 fold change and adjusted p-values.
- Protein Modeling – The Cryo-EM structure of SLC4A4 protein (pdb id: 6caa) was modeled by SWISSMODEL.
- Virtual Screening and Docking – The modeled protein was subjected to docking with a library of FDA-approved drugs obtained from DrugBank (via DrugRep) using AutoDock Vina and redocked with CB-DOCK2.
- Molecular Dynamics Simulation – The lead drug candidate (Nilotinib) was further validated for its binding stability using a 50-ns MD simulation with GROMACS, with analysis of RMSD, RMSF, hydrogen bonding, radius of gyration, and SASA metrics.
As this study is observational and computational in nature, it did not involve any live or human subjects. All datasets were retrieved from publicly accessible databases, and the analytical tools employed are open-source or web-based. Such an approach promotes transparency and ensures the reproducibility of computational drug repurposing research.
3. RESULTS
3.1. Expression of SLC4A4 from Expression Atlas Datasets
Analysis of SLC4A4 Expression from Expression Atlas Datasets. To investigate SLC4A4 expression in colorectal cancer (CRC), we performed a comprehensive analysis using RNA-sequencing (RNA-seq) and array-based data from the Expression Atlas database. When the analysis was restricted to datasets with CRC, significant differences in SLC4A4 were observed. The SLC4A4 expression was downregulated consistently in these datasets. It was identified with the thresholds of Log2 Fold Change (log2FC)<-1 and an adjusted p-value (adj. p-val) ≤0.05. This downregulation of SLC4A4 was observed in many CRC datasets and is shown in Table 1, suggesting a possible role for SLC4A4 in CRC development.
3.2. Molecular Modelling
A 3D model of SLC4A4 with template Q9Y6R1 was built as provided in Table 2. The model was analysed using the Protein Structure and Model Quality Assessment Tool of Swiss-Model Server, which resulted in favorable scores: GME (Global Model Quality Estimate) = 0.71; Ramachandran Favoured (%) = 90.69. Furthermore, the MolProbity Score was 1.64 (Table 3) [32]. These evaluations highlighted the feasibility of employing this model for further structure-based 6CAA analysis and design.
| Gene | Species | Experiment Accession | Comparison | log_2 fold Change | Adjusted p-value |
|---|---|---|---|---|---|
| ENSG00000080493 | homo sapiens | E-GEOD-19249 | 'Colon cancer' vs 'normal' in 'colon; Fresh-frozen tissue' | -5.9 | 2.18433988997483E-05 |
| ENSG00000080493 | homo sapiens | E-GEOD-76987 | 'Colon adenocarcinoma' vs 'normal' | -4.4 | 5.53014902194911E-40 |
| ENSG00000080493 | homo sapiens | E-GEOD-68086 | 'Colorectal carcinoma' vs 'normal' | -3.2 | 2.94812359574088E-09 |
| ENSG00000080493 | homo sapiens | E-GEOD-19249 | 'Colon cancer' vs 'normal' in 'colon; Formalin-fixed paraffin-embedded tissue' | -3.1 | 0.02006192248942 |
| ENSG00000080493 | homo sapiens | E-GEOD-4183 | 'Colon adenoma' vs 'none' | -3 | 6.23893701946427E-05 |
| ENSG00000080493 | homo sapiens | E-GEOD-50760 | 'Colorectal cancer metastatic in the liver' vs 'normal' | -2.5 | 2.04883548287025E-06 |
| ENSG00000080493 | homo sapiens | E-GEOD-4183 | 'Colorectal cancer' vs 'none' | -2.2 | 0.049104604966632 |
| Template | Seq Identity | Oligo-state | Found By | Method | Resolution | Seq. Similarity | Range | Coverage | Description |
|---|---|---|---|---|---|---|---|---|---|
| Q9Y6R1.1.A | 99.5 | Monomer | AFDB search | AlphaFold v2 | --- | 0.61 | 35 - 1035 | 0.97 | Electrogenic sodium bicarbonate cotransporter 1 |
| Model | Built With | Oligo-State | Ligands | GMQE | MolProbity Score | Ramachandran Favoured |
|---|---|---|---|---|---|---|
| 1 | ProMod3 3.4.0 | Monomer | None | 0.71 | 1.64 | 90.69% |
3.3. Molecular Docking
Drug repurposing for approved drugs was done with a DrugRep tool running with default parameters, considering compounds in DrugBank (version 5.1.7). The findings from virtual screening and docking were the identification of a key binding pocket in the protein SLC4A4, which was the major ligand-binding site. The features of this pocket are summarized in Table 4. The pocket (Pocket 1) is a substantial 3676 Å3 at (−1.4, −18.4, −0.4) in the xyz space. It contains critical amino acid residues, including hydrophilic residues such as Aspartic Acid (ASP), Glutamic Acid (GLU) , and Lysine (LYS), and hydrophobic residues such as Phenylalanine (PHE) and Leucine (LEU). This mixed composition means that the pocket is diverse enough to bind multiple ligands via various types of interactions, which is imperative to understand, as many of the best ligands for our four target drugs are characterized by highly positive binding (which is in our favor).
Redocking was carried out with selected compounds using CBDOCK2 to obtain the highest-ranked docking pose for each compound, based on the scoring functions: DB01396 (Digitoxin), DB00762 (Irinotecan), and DB04868 (Nilotinib). Interestingly, Nilotinib had the best docking score, suggesting a higher binding affinity (Table 6). The top three hits, identified based on cavity-based docking, are listed in Table 5.
| Volume | Center | Size | |
|---|---|---|---|
| 3676 | -1.4,-18.4,-0.4 | 22,21,19 | Chain A (ASP960 ASN763 ASP966 VAL760 VAL961 ASP947 GLY387 PHE531 LYS974 PHE536 VAL533 PRO963 GLN952 LEU392 GLU971 LYS970 HSD386 LEU955 ALA773 ASP959 LYS765 GLU766 SER956 LYS768 LYS944 TYR775 ARG764 LEU769 GLU541 CYS389 ILE962 ARG943 GLN534 ARG538 GLY772 LYS967 TYR535 GLU964 GLU391 VAL940 THR537 ASP388) |
| DrugBank ID | Docking Score (kcal/mol) |
|---|---|
| DB01396 | -10.2 |
| DB00762 | -10 |
| DB04868 | -9.6 |
| FDA-approved Drugs | Pocket ID | Vina Score (kcal/mol) | Cavity Volume (Å3) | Center (x, y, z) | Docking Size (x, y, z) | Pocket Seq. |
|---|---|---|---|---|---|---|
| DB04868 (Nilotinib) | C1 | -11.2 | 3676 | -1, -18, 0 | 24, 24, 24 | Chain A: HIS386 GLY387 ASP388 CYS389 GLU391 LEU392 GLN534 PHE536 THR537 ARG538 GLU541 VAL760 ASN763 ARG764 LYS765 GLU766 LYS768 TYR775 GLN877 PHE895 ILE896 TYR897 HIS900 ARG943 LYS944 GLN952 HIS953 LEU955 SER956 PHE957 ASP959 ASP960 VAL961 ILE962 PRO963 GLU964 LYS965 LYS967 LYS968 GLU971 |
| DB00762 (Irinotecan) | C3 | -10 | 2519 | -9, 4, -1 | 30, 30, 30 | Chain A: LYS233 ASN422 ILE423 GLN424 SER427 LYS670 SER672 PRO673 PHE675 THR677 THR678 ARG680 LYS681 LEU682 ASP685 PHE686 GLU766 HIS767 ASP809 LYS812 MET813 GLU814 THR815 GLU816 THR817 SER818 ALA819 PRO820 GLY821 GLU822 GLN823 PRO824 PHE826 ARG830 GLY875 |
| DB01396 (Digitoxin) | C4 | -9.8 | 2046 | -23, 2, 27 | 33, 33, 33 | Chain A: TRP87 ILE88 LYS89 PHE90 LYS93 VAL94 GLN96 SER102 HIS105 VAL106 ALA107 THR108 LEU109 SER110 LEU111 LYS291 HIS294 GLU295 ARG298 THR302 SER305 ASP306 GLU307 GLU328 PHE329 GLU332 VAL333 ILE334 VAL335 LEU336 PRO338 TRP341 ASP342 PRO343 ALA344 ARG346 ILE347 GLU348 PRO349 PRO350 LYS351 SER352 LEU353 PRO354 SER355 SER356 ASP357 LYS358 ARG359 |
The docking complex between the modeled 6CAA protein and Nilotinib (Fig. 1) was then used for subsequent molecular dynamics simulation was also performed to assess its stability and binding properties.
Protein 6CAA and nilotinib docking complex.
3.4. Molecular Dynamics Simulation
For the protein–nilotinib complex, the system showed stable dynamics, and the protein reached a maximum RMSD of ~1.5 nm over a 50-ns simulation (black line in Fig. 2A). The ligand also maintained fluctuations at ~0.30 nm RMSD over the same period (red line in Fig. 2A), indicating a stable positioning within the binding pocket. In general, the RMSD results suggest that the ligand remained stably bound to the protein's active site.
The RMSF analysis showed that the N-terminal part of the protein fluctuated slightly at about 1.0 nm and that the C-terminal was more flexible with its fluctuations up to ~2.0 nm as represented by the black line in the Fig. (2B). In contrast, the ligand was much less flexible, staying within ~0.5 nm during the whole simulation (the red line).
The radius of gyration(Rg) results showed ligand (red line) fluctuated up to 0.5 nm while protein(black line) showed a slow decreasing from 4.0 nm to about 3.8 nm at the end of the simulation. This indicates that the ligand is stably located in the binding site of the protein (Fig. 2C).
Hydrogen bonding analysis showed that the ligand established up to seven hydrogen bonds intermittently at 4, 8, 9, 11, 12, 14, and 16ns during the simulation. At the end of the 50-ns run, three hydrogen bonds survived within the protein-ligand complex, as shown in Fig. (2D).
The Solvent Accessible Surface Area (SASA) for the protein only (black line) and the protein-ligand complex (red line) during the entire MD simulation. The two graphs superimpose closely, indicating little fluctuation, and demonstrating that the system was stable over the entire simulation period (Fig. 2E).
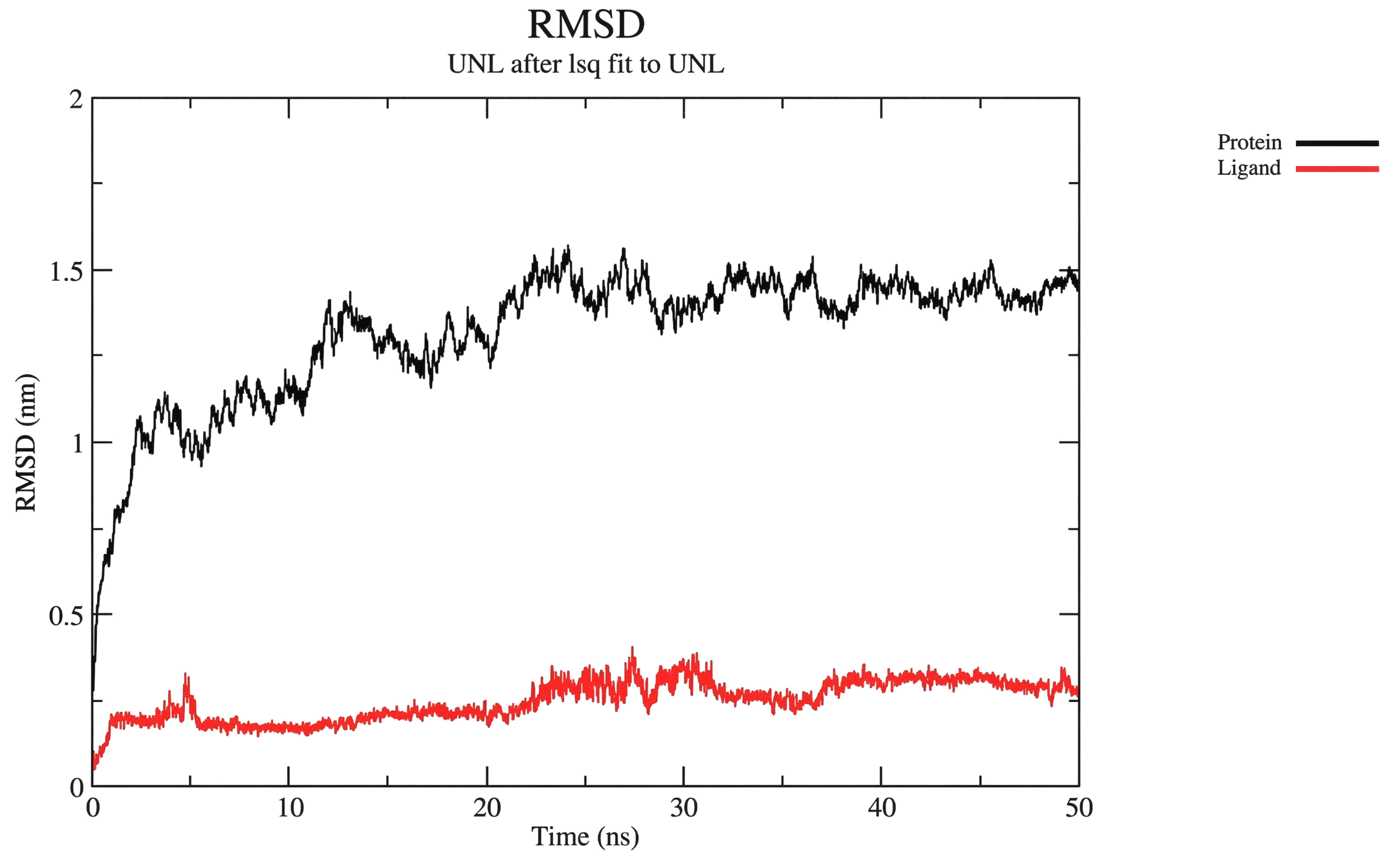
Protein-ligand RMSD.
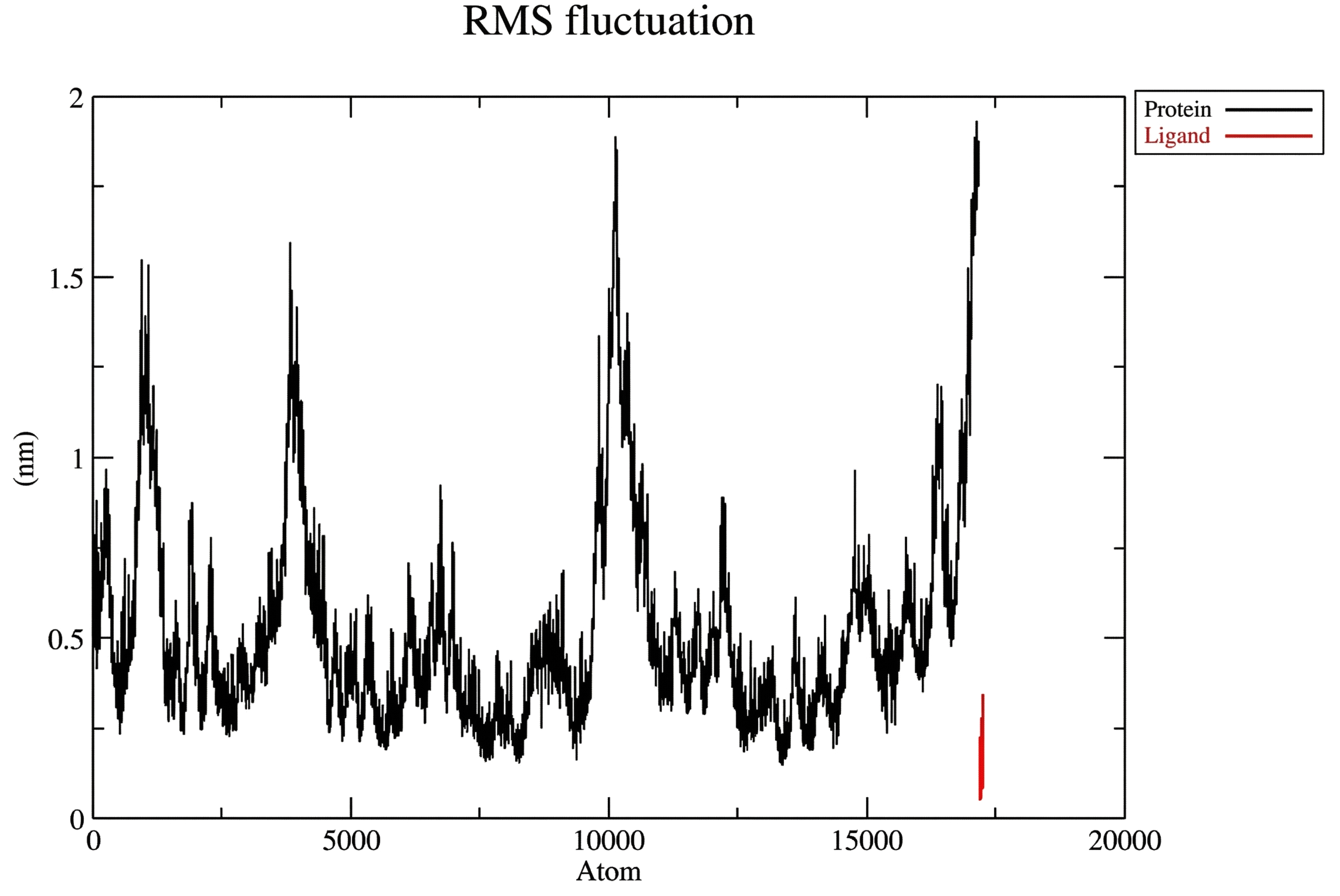
Protein-ligand RMSF.
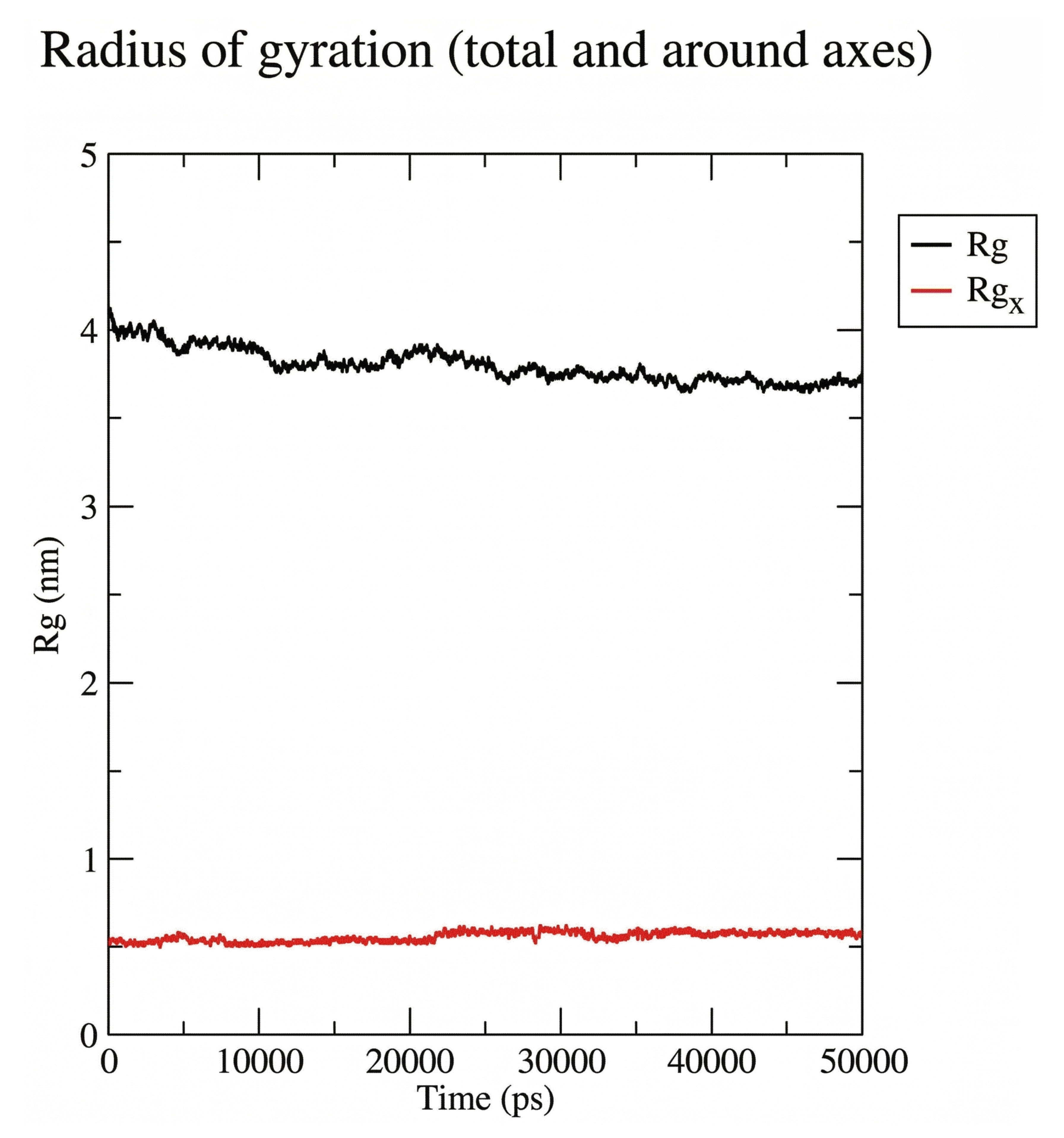
Protein-ligand radius of gyration.
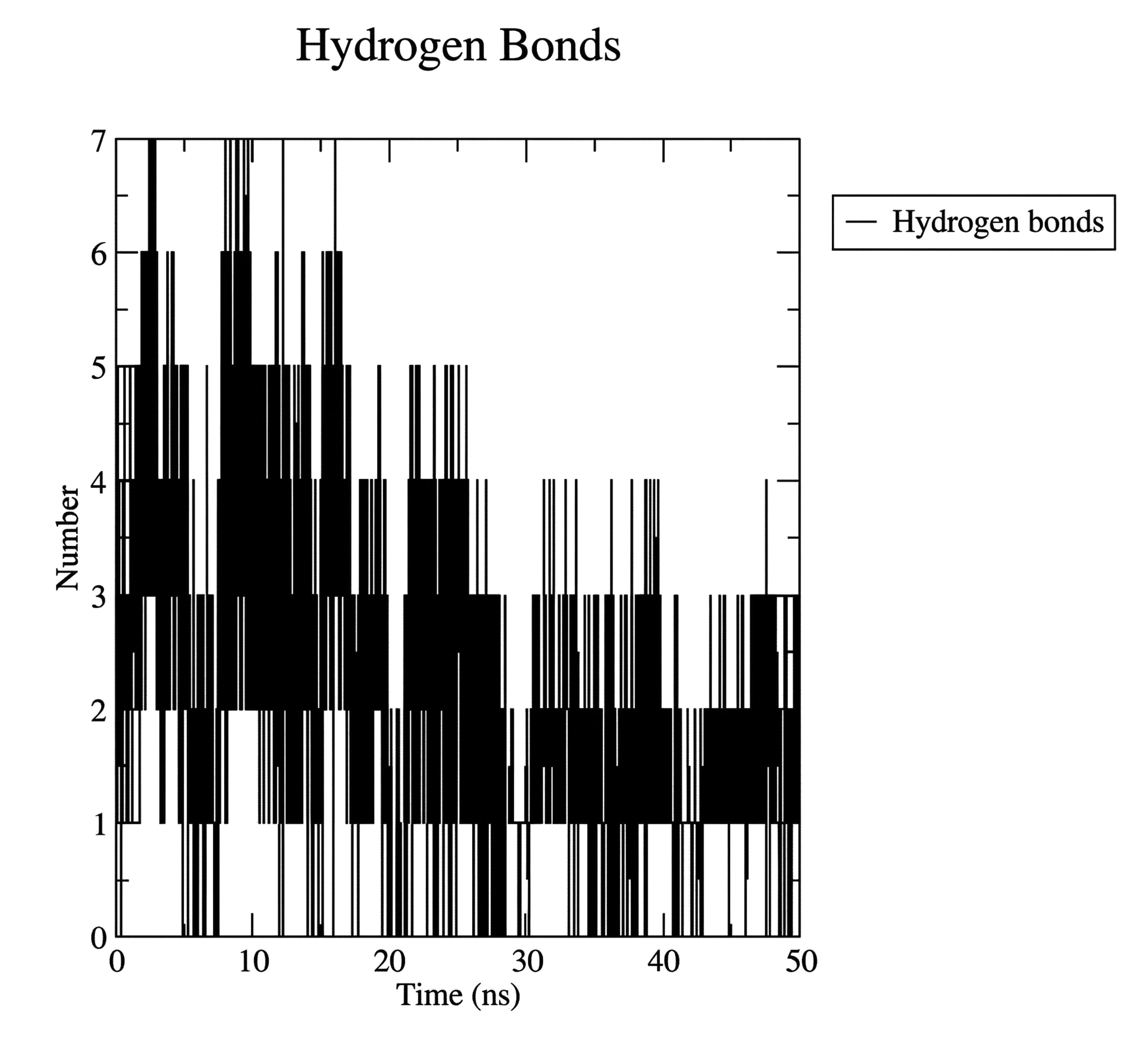
Protein-ligand hydrogen bond interaction.
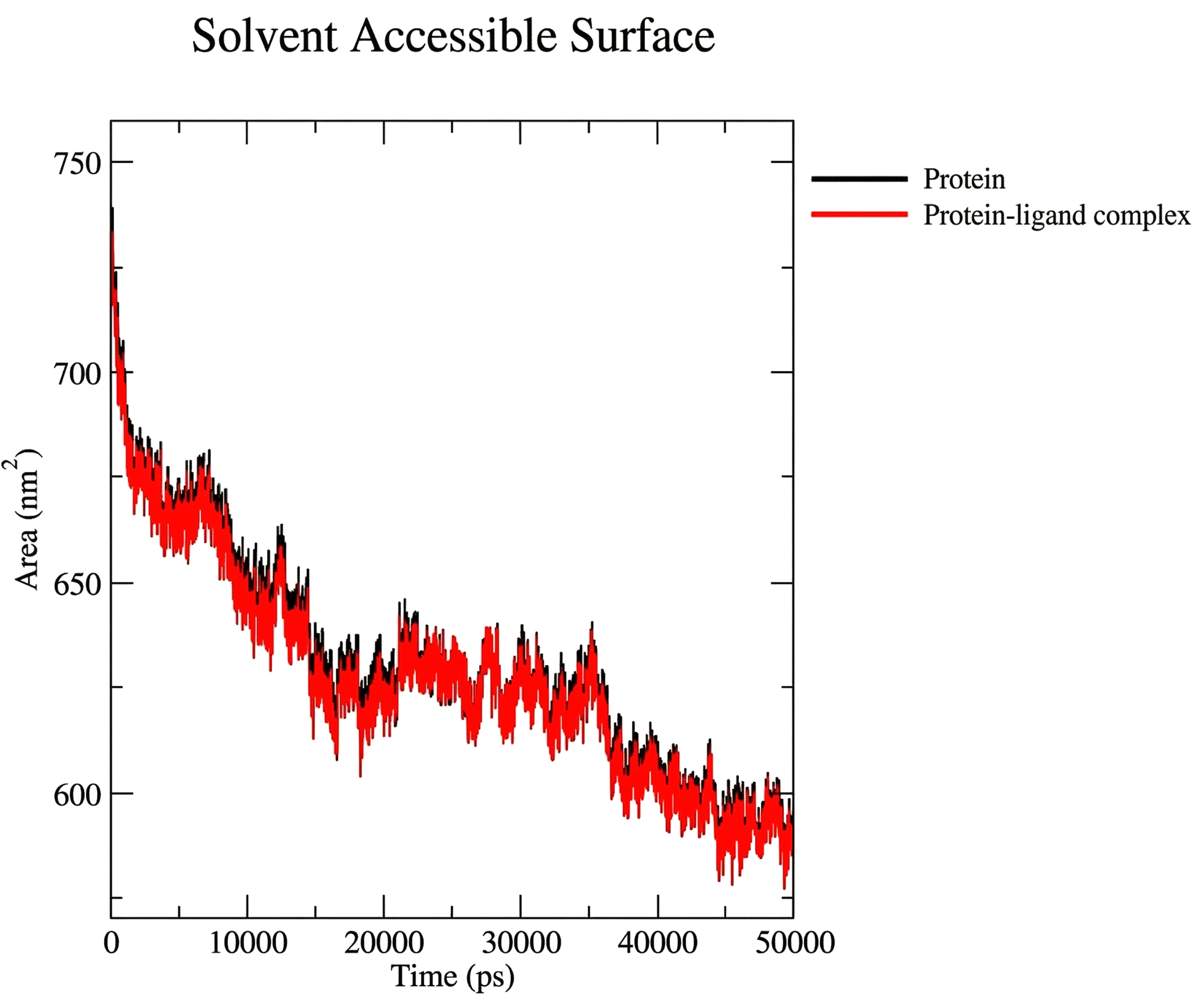
Protein-ligand SASA analysis.
Upon energy minimization, the Nilotinib–protein complex showed a number of significant bonds. At the initial conformation (0 ns), the compound makes an H-bond with Ser38, Arg40, Pro105, Ile132, Gln107, Ser349, and Glu351. Pi-interactions were observed with residues such as Arg40, Ala104, Lys277, Pro279, His351, Thr861, Ser862, Ala863, and Lys869. Additionally, van der Waals interactions involved Val35, Lys37, Tyr39, Arg41, Pro106, Leu108, Lys133, Met348, Thr859, and Glu861 (Fig. 3A).
At the 25 ns mark, Nilotinib still had four hydrogen bonds with Cys38 and Glu351. Pi-interactions were observed with Lys37, Ser38, and His354, and van der Waals interactions with Pro36, Tyr39, Arg40, Ala104, Pro105, Pro106, Gln107, Leu108, Ser349, and Asp350 (Fig. 3B).
At 50 ns (end of simulation), the ligand was hydrogen-bonded to Ser38 and Ser349. Pi-interactions with Pro36, Ala104, Pro105 and His354 and van der Waals interactions with Tyr39, Arg40, Arg41, Pro106, Gln107, Leu108, Glu351, and Lys814 (Fig. 3C).
From 0 ns to 50 ns, Nilotinib remained buried in the protein's binding pocket, suggesting stable binding with diverse favorable interactions throughout the simulation.
4. DISCUSSION
The importance of SLC4A4 in cellular pH homeostasis is undisputed, and its biological relevance in colorectal cancer is far more than the mere maintenance of pH balance. A few recent reports have demonstrated the importance of this molecule in influencing the tumor microenvironment in relation to EMT and metastasis. Besides having direct impacts on cancer cells, SLC4A4 is important for the inflammatory TME. SLC4A4 expression has been positively associated with infiltration of several immune cell types, including CD8+ T cells, dendritic cells, macrophages, and NK cells, in colorectal cancer (CRC) tissues [18, 31]. This is meaningful because these are the key immune cells required for an efficient anti-tumor immune response. Thus, decreased SLC4A4 can induce a TME that is less immunostimulatory and more immunosuppressive, allowing nasal tumors to evade immune surveillance and thrive. This observation indicates that restoring or maintaining SLC4A4 activity might represent a new avenue for “re-educating” the TME to become more unfavorable to cancer cells and more conducive to immunotherapy. In addition, studies have suggested that higher SLC4A4 expression is associated with better response to some immunotherapies, such as Nivolumab and Ipilimumab, whereas lower expression is associated with better response to other targeted therapies [31]. This dual functional attribute makes SLC4A4 a promising target as a therapeutic agent and as a predictive biomarker for treatment options and personalized medicine.
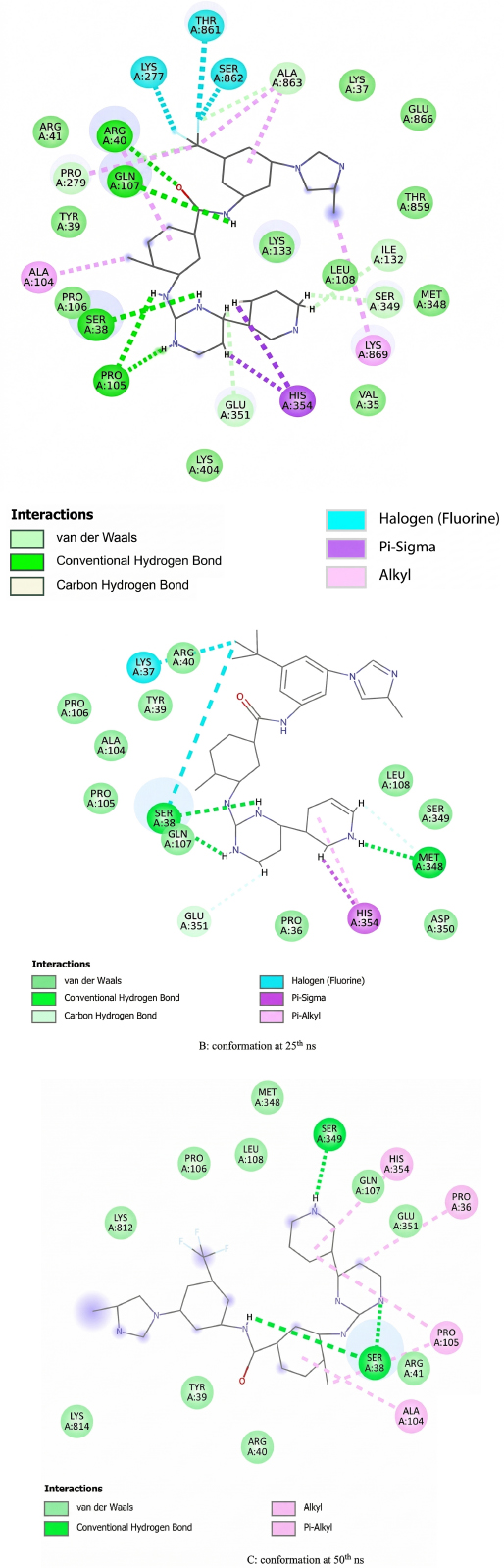
Protein binding site conformations at different timepoints; (A): conformation at 0th ns. (B): conformation at 25th ns. (C): conformation at 50th ns.
The biological relevance of SLC4A4 is further supported by its involvement in Epithelial-Mesenchymal Transition (EMT), a crucial step in cancer metastasis. EMT is a multistep cellular program in which epithelial cells lose cell–cell adhesion and acquire migratory and invasive abilities, allowing them to detach from the primary tumor and metastasize to distant organs. Evidence shows that a reduced SLC4A4 expression was associated with higher lymph node and distant metastasis and modulated partial EMT phenotypes required for cancer cell migration and invasion. These results indicate that SLC4A4 functions as a tumor suppressor by repressing EMT. In so reactivating SLC4A4, it might be feasible to block a critical metastatic circuit, ultimately preventing Going beyond CRC and boosting outcomes for patients [17]. That adds one potent layer to why one would want to target SLC4A4, since it is the definition of a most deadly part of cancer progress.
The decreased expression of SLC4A4 in CRC specimens compared with normal tissues indicates that this gene may have an important function in the development of CRC [40]. SLC4A4 encodes a bicarbonate ion transporter protein essential for pH regulation and cell homeostasis [41]. Aberrant expression of this gene has been associated with many cancers, including CRC, signifying a potential therapeutic role [42, 43].
Repurposing of drugs, which means finding new indications for known drugs, is a time/test resource-saving strategy in drug discovery. In this respect, SLC4A4 targeting may be a novel therapeutic approach. Discovery of compounds that block SLC4A4 might allow for reestablishment of normal cellular physiology and antagonize tumor growth [19]. In addition, disruption of the tumor microenvironment through SLC4A4 inhibition could sensitize other therapies as well, providing a new avenue for combination therapy.
Nilotinib is a second-generation tyrosine kinase inhibitor, designed as a more potent and selective inhibitor against the BCR-ABL fusion protein and the first-line treatment for chronic myeloid leukemia (CML). Nevertheless, growing preclinical evidence indicates that nilotinib exerts broader anticancer activity and has potential for repurposing in other cancers, such as colorectal cancer (CRC) [44].
Nilotinib has also been shown to block the activity of a number of other tyrosine kinases involved in cancer. Interestingly, DDR1, as for BCR-ABL in chronic myelogenous leukemia, has also recently been shown to be an important target of nilotinib in CRC. DDR1 mediates tumor cell invasion and metastasis through downstream effectors, including β-catenin signaling. Treatment with nilotinib in cell culture and animal CRC models has demonstrated robust inhibition of CRC invasion and metastatic dissemination by inhibiting DDR1 kinase activity and impeding continuous DDR1 signaling [45, 46].
In addition to its direct tumoricidal effects, nilotinib has immunomodulatory effects that might improve its activity in CRC [18]. Dong et al. (2024) revealed the potential of the tyrosine kinase inhibitor nilotinib in restoring Major Histocompatibility Complex class I (MHC-I) expression in CRC cells, leading to increased tumor immunogenicity and sensitizing to immune checkpoint inhibitors, i.e., anti-PD-L1 therapy. This dual mechanism of action direct inhibition of tumor-promoting kinases and potentiation of anti-tumor immunity makes nilotinib a particularly promising agent for the development of combination treatment strategies in CRC [47].
Notably, the clinical safety and pharmacokinetics of nilotinib have been well characterized following extensive use in patients with chronic myeloid leukemia. Long-term clinical research has revealed that nilotinib is overall well-tolerated, with manageable adverse events and a predictable pharmacological profile [48]. These properties make it suitable for rapid clinical translation and repurposing in CRC, given that dosing regimens and safety monitoring guidelines are well established [49].
In conclusion, the established efficacy of nilotinib in targeting DDR1-mediated signaling, its contribution to enhancing anti-tumor immunity, and its well-characterized safety make it an attractive candidate for further studies as a therapeutic agent in CRC. To the best of our knowledge, this is the first in silico drug repositioning study leveraging FDA-approved drugs to predict SLC4A4 inhibitors in colorectal cancer. Although SLC4A4 has been implicated in pH regulation and its prognosis has been described, none of the aforementioned studies evaluated FDA drugs for direct targeting of the transporter, which makes this work a first exploratory step in the CRC drug discovery.
5. LIMITATIONS
However, several caveats should be kept in mind. First, this study is fully computational and has not been experimentally validated. Therefore, the results should be considered a starting point for subsequent work. The predictions may be biased by the off-targets' full landscape and by the limitations of the protein modelling and docking protocols to fully describe the dynamic nature of protein-ligand interactions in a cellular environment. In addition, we realize that we could not add external control ligands or known inhibitors for benchmarking, as the DrugRep server selected for this work does not allow users to add their own ligands but only allows screening its curated FDA-approved drug library. In addition, no comparison with a deduced SLC4A4 inhibitor was possible, as there are no inhibitors reported at the time of writing. Similarly, binding free energy (MMPBSA or MM-GBSA) calculations were not performed, but would be valuable from an energetic perspective. These will be pursued in future studies along with experimental bioassays. Future work using programmable docking platforms may overcome this by providing particular controls for better benchmarking. In summary, future studies should include experimental validation and investigation of combinational therapy to fully exploit the therapeutic potential of nilotinib and other prospective SLC4A4- targeting agents in CRC.
CONCLUSION
In conclusion, the docking and molecular dynamics simulations of nilotinib with the modeled 6CAA protein have yielded several significant findings, including high affinity and stability, consistent binding poses, protein flexibility, strong hydrogen bonding, and stable conformation, as mentioned in the results section. The conformational analysis of the MD simulation results confirmed the presence of nilotinib within the binding site region of the protein. Considering these findings, nilotinib emerges as a promising candidate for further investigation and potential development due to its high affinity for binding strongly and stably without significantly distorting the protein conformation. Further research is necessary to validate the effectiveness of nilotinib in colorectal cancer (CRC) and to elucidate the precise mechanisms by which it interacts with SLC4A4 and other relevant molecular targets. Nevertheless, the repurposing of nilotinib for CRC treatment holds significant promise and could lead to more effective and personalized therapeutic strategies.
DECLARATION
All figures (docking images, MD plots, structural models) and all tables were generated by the tools like: Expression Atlas public datasets, SWISS-MODEL, DrugRep, CB-DOCK2, GROMACS, PyMOL, Grace plotting softwares.
AUTHORS’ CONTRIBUTIONS
The authors confirm contribution to the paper as follows: K.P., P.P.G., R.R.K.N.: Conceptualization, methodology, formal analysis, investigation, resources, data curation, writing, draft preparation; S.L.K., P.S.S., R.R.K.N.: Supervision. All authors have read and agreed to the published version of the manuscript.
LIST OF ABBREVIATIONS
| CRC | = Colorectal Cancer |
| SLC4A4 | = Solute Carrier Family 4 Member 4 |
| pH | = Potential of Hydrogen (measure of acidity/basicity) |
| p-EMT | = Partial Epithelial–Mesenchymal Transition |
| MHC-I | = Major Histocompatibility Complex Class I |
| PD-L1 | = Programmed Death-Ligand 1 |
| MD | = Molecular Dynamics |
| RMSD | = Root Mean Square Deviation |
| RMSF | = Root Mean Square Fluctuation |
| Rg | = Radius of Gyration |
| SASA | = Solvent Accessible Surface Area |
| PME | = Particle Mesh Ewald (method for computing long-range electrostatics) |
| TIP3P | = Transferable Intermolecular Potential with 3 Points (water model) |
| NVT | = Constant Number of Particles, Volume, and Temperature |
| NPT | = Constant Number of Particles, Pressure, and Temperature |
AVAILABILITY OF DATA AND MATERIALS
The data supporting the findings of the article are available in the Expression Atlas Repository at https://www.ebi.ac.uk/gxa/home, under publicly accessible experiment accession IDs listed in Table 1 of the manuscript (e.g., E-GEOD-19249, EGEOD-76987, E-GEOD-68086).
ACKNOWLEDGEMENTS
Declared none.


