Artificial Intelligence in PubMed-indexed Biomedical Research: Results and Analysis
Abstract
Introduction
Artificial intelligence (AI) has rapidly emerged as a transformative force in biomedical research, driving advances in data interpretation, diagnostics, and therapeutic strategies. The objective of this study was to conduct a large-scale bibliometric analysis of AI-related biomedical publications indexed in PubMed between 1950 and 2024, with the aim of identifying temporal trends, global research contributions, and thematic focus areas.
Methods
A retrospective bibliometric analysis was performed on 76,722 PubMed-indexed articles. Data were retrieved using a Python-based pipeline integrating National Center for Biotechnology Information (NCBI) Entrez Programming Utilities and Biopython. Metadata-including PubMed Identifier (PMID), title, abstract, authors, publication date, journal, Medical Subject Headings (MeSH) terms, Digital Object Identifier (DOI), and country of origin-were extracted, cleaned, and compiled into a structured dataset. Articles were analyzed for completeness, publication patterns, geographic distribution, journal outlets, and thematic focus using MeSH keywords.
Results
AI-related publications remained scarce until 2015, after which output expanded exponentially, reaching 20,135 articles in 2024. The United States contributed the largest share (32.5%, n = 24,958), followed by England (22.5%) and Switzerland (n = 13,936). Thematic analysis revealed machine learning (n = 16,281), deep learning (n = 12,297), and neural networks (n = 7,117) as dominant areas, while AI ethics appeared in only 110 publications. Metadata completeness varied, with notable gaps in MeSH indexing (41.6%), abstracts (9%), and Digital Object Identifiers (DOI) (2.6%). Research was widely disseminated across more than 6,000 journals, with Sensors, Scientific Reports, and Public Library of Science ONE (PLOS ONE) as leading outlets.
Discussion
The findings highlight AI’s transition from a peripheral topic to a core pillar of biomedical science, with rapid growth driven by technological advances and global health demands. Despite widespread adoption across multiple disciplines, gaps remain in ethical engagement and equitable global representation. Metadata inconsistencies also pose challenges for systematic synthesis and bibliometric analyses.
Conclusion
AI has become a central component of biomedical research, characterized by exponential growth, interdisciplinary adoption, and global expansion. However, limited attention to AI ethics and persistent disparities in research representation underscore the need for targeted policy, funding, and governance strategies. Continued bibliometric monitoring is essential to ensure responsible and inclusive integration of AI into clinical and research practice.
1. INTRODUCTION
Technological interventions in biomedical science have not followed a singular or linear path. Systems once designed for narrow tasks now process clinical language, decode images, and adapt to unstructured datasets with surprising fluidity [1, 2]. Not all of this is new; predictive modeling and language parsing were present in earlier forms, yet the capacity to scale and learn from vast biomedical records has only recently taken hold. What this indicates is not just methodological refinement, but a structural shift in how knowledge is generated and operationalized across domains.
Much of the recent intensity in biomedical innovation owes itself to a convergence of factors: expanding digital repositories, increased cloud access, and widely shared algorithmic frameworks. These conditions have sparked a diversification of applications-from diagnostic imaging and intensive care modeling to genomics-driven therapy selection [3, 4]. However, it is not the tools themselves that demand attention, but the velocity and volume with which they have been adopted across traditionally siloed medical fields.
Artificial intelligence has gained more and more significance in the healthcare sector because of its capability to analyze large volumes of data, recognize trends, and create new insights that can be used to improve decision-making. Its application is useful in various ways. AI-based imaging and pathology systems facilitate earlier and more precise diagnosis of diseases, machine-learning-based treatment planning methods focus on individualized medicine and adjust therapies to genetic and clinical backgrounds, and operational management applications use AI to streamline processes, resource distribution, and patient scheduling. Automated processing of electronic health records is assisted through natural language processing to enhance documentation and clinical communication. Outside of clinical services, AI has applications in population health monitoring, disease outbreak forecasting, and in managing health data databases of scale. All of them, together, not only enhance efficiency and accuracy but also allow saving money, reducing errors, and broadening access to high-quality care, which proves the importance of AI in the modern healthcare industry [3-6].
Literature output offers a distinct lens into this transformation. Databases such as PubMed, though often perceived as static archives, now reflect an evolving thematic and geographic distribution of work related to intelligent systems in medicine. Still, growth alone is not the only metric of importance. Questions remain about how this research is framed, who contributes to it, and which aspects of healthcare remain unexamined [5, 6].
There is no universally agreed-upon method for mapping these dynamics. Yet, bibliometric analysis stands as a robust, if imperfect, means of tracing conceptual, temporal, and geographic patterns. By analyzing metadata from thousands of indexed records, one can identify trends in authorship, topical saturation, and systemic omissions. These findings are often as valuable to policy and institutional strategy as they are to scholarly assessment [7, 8].
This current investigation does not attempt to answer every question, but rather to chart the contours of a rapidly maturing research frontier.
It is worth mentioning that PubMed was launched in January 1996 by the National Center for Biotechnology Information (NCBI) at the U.S. National Library of Medicine (NLM). It was made publicly available in June 1997. It provides access to the Medical Literature Analysis and Retrieval System Online (MEDLINE) database, which dates back to the 1960s and indexes biomedical literature from as early as 1946.
Drawing on indexed records from 1950 to 2025, it examines publication rates, country-level activity, journal outlets, and metadata integrity to offer a fragmented yet meaningful view of how computational systems have insinuated themselves into the logics and practices of biomedical inquiry.
2. METHODOLOGY
2.1. Study Design and Data Source
This investigation, though rooted in bibliometric principles, adopted a historical lens to observe evolving research practices within the biomedical field. The study’s architecture was underpinned by a custom-built Python script, crafted specifically to handle automated interactions with NCBI Entrez E-utilities via Biopython (version 1.79+). This design wasn’t simply about automation; it allowed for consistent metadata structuring, supporting retrieval efforts spanning multiple decades. The reproducibility this enabled was not incidental, but rather necessary given the scope and scale of the literature involved. Parsing, query segmentation, and batch retrieval were handled programmatically, ensuring the research maintained integrity over time.
Interestingly, the decision to use PubMed as the central corpus stemmed not only from its breadth but from its structural assets. The Medical Subject Headings (MeSH) framework-integrated directly into PubMed indexing-played a crucial role in enabling topical cohesion during the parsing phase. Beyond just standardized classification, MeSH terms provided a stable thematic scaffold. Although other repositories exist, few offer the same granularity or temporal traceability. PubMed, thus, emerged not merely as a data source, but as an analytical foundation capable of reflecting complex temporal and geographic shifts in knowledge generation [5, 9].
2.2. Search Strategy
Query segmentation wasn’t merely a methodological necessity. It evolved as a strategic decision born from the dual pressures of temporal comprehensiveness and technical restraint. The dataset’s architecture was informed by the uneven distribution of publications across time, particularly during the initial decades (1946–1994), when output relevant to intelligent computational methods remained infrequent. For such intervals, broad temporal windows were acceptable. However, this leniency did not extend into the post-2008 publication surge, which required annual granularity and later month-by-month scrutiny.
Rather than a single continuous sweep, the extraction routine was fragmented. The search syntax-targeting the Title/Abstract field with the query term “artificial intelligence”-was framed using PDAT (Publication Date) parameters to anchor each segment. To enhance transparency and reproducibility, we explicitly defined the search terms applied. The core query used was “artificial intelligence”[Title/Abstract], with supplementary terms such as “machine learning,” “deep learning,” “neural networks,” “natural language processing,” and “radiomics” tested to confirm retrieval sensitivity and thematic coverage. This ensured that the dataset comprehensively reflected AI-related biomedical research indexed in PubMed.
Consistency in this formulation, such as “artificial intelligence”[Title/Abstract] AND (“start_date”[PDAT]: “end_date”[PDAT]), allowed the framework to remain stable even as frequency intensified. Data calls to Entrez.esearch were limited to 1,000 records per batch, and inter-query delays were intentionally programmed not just to conform to Application Programming Interface (API) etiquette but also to preserve response stability across retries. Scalability and error resilience weren’t afterthoughts. They were embedded by design, driven by lessons learned from prior interruptions and latency issues in literature mining.
2.3. Data Extraction and Processing
The metadata phase didn’t follow immediately in a linear fashion-it emerged as a necessary layer once the PubMed Identifiers (PMIDs) had been gathered. Using Entrez.efetch with rettype=”medline” and retmode=”text”, the system retrieved full records in a consistent, readable format. Only then could parsing begin, with Bio.Medline.parse from Biopython performing the heavy lifting-not as an auxiliary step, but as the pivot point for structured bibliometric interpretation.
Parsing was not limited to the obvious. Yes, article titles and abstracts were included-fundamental for thematic indexing-but the process reached further. All records were programmatically retrieved through the NCBI Entrez E-utilities using Biopython (version 1.79+). Each request was limited to 1,000 records per batch, with programmed inter-query delays to ensure compliance with API requirements and prevent retrieval errors. The metadata fields extracted included PMID, title, abstract, author list, journal, publication type, MeSH terms, DOI, affiliations, and country of origin.
The PubMed ID (PMID) served as the anchor, while journal names added disciplinary context. Dates, of course, were extracted-not merely for cataloging but to anchor time-series evaluations. In parallel, publication type classifications such as “Journal Article” or “Review” provided signals about the depth and orientation of the scientific discourse.
Less visible fields proved equally crucial. MeSH descriptors enabled standardized thematic alignment, while author affiliations pointed not just to institutions, but to broader geopolitical patterns in the evolution of computational health studies. DOIs, retrieved from the Locator Identifier (LID) field, were less glamorous but indispensable for persistent referencing. The ‘country’ tag-frequently overlooked-played a quiet yet pivotal role in geographic trend analysis.
Eventually, everything converged into a structured Comma-Separated Values (CSV) repository not just for storage, but for all downstream inferential and descriptive routines.
2.4. Data Cleaning and Validation
Once the initial set of PMIDs had been assembled, attention shifted to the more nuanced task of metadata extraction. Rather than beginning with thematic analysis, the process first required careful structuring of bibliographic records. The Entrez.efetch utility-configured with the parameters rettype=”medline” and retmode=”text”-served as the conduit for retrieving entries in a format that supported consistent parsing and interpretation.
Parsing was not undertaken arbitrarily. The Biopython module, particularly the Bio.Medline.parse function was leveraged at this stage, allowing the retrieval process to unfold through a controlled pipeline capable of dissecting structured MEDLINE entries with minimal data loss. Cleaning procedures included deduplication of records, standardization of author names using a semicolon delimiter, cross-checking missing abstracts and DOIs, and validation of country fields for geographic analysis. These steps ensured a consistent and reproducible dataset for downstream bibliometric analysis.
Interestingly, it was not just the volume of records, but the variability across fields-titles, abstracts, and DOIs-that introduced complexity into the extraction procedure.
Crucially, each entry yielded multiple elements-some expected, others surprisingly inconsistent across records. Article identifiers (PMIDs), titles, and abstracts were prioritized for their central role in guiding later thematic segmentation. In parallel, author lists were standardized using a semicolon delimiter, which proved helpful when mapping collaborative networks. Journal names, less frequently used in preliminary analytics, were nonetheless retained for evaluating publication venues and disciplinary reach.
The publication date for each article was captured not as a timestamp, but as a temporal anchor for mapping shifts in output over defined eras. DOIs-often buried in the LID field-were parsed to ensure stable linkage for referencing purposes. Additionally, classification into publication types, whether as original research, reviews, or other forms, was maintained, recognizing that the structure of scholarly contributions could influence citation dynamics and dissemination pathways.
3. RESULTS
A total of 76,722 unique publications related to artificial intelligence (AI) in biomedical research were analyzed in this study, each indexed in PubMed and accompanied by metadata fields including PMID, title, abstract, author list, journal name, publication date, DOI, publication type, MeSH terms, affiliations, and country of origin. This comprehensive dataset enabled an in-depth exploration of publication patterns, geographical reach, thematic trends, and research dissemination across time and disciplines.
3.1. Data Completeness
The overall quality of the dataset was high, but some disparities in metadata coverage emerged. Abstracts were absent in approximately 9% of articles (n = 6,912), often in older or brief report formats. MeSH term coverage was significantly lower, missing in 41.6% of entries (n = 31,902), particularly in recent publications that had not yet undergone full indexing. DOI identifiers were lacking in about 2.6% of records (n = 2,021), again predominantly among older studies. In contrast, fields such as PMID, publication type, date, and country were consistently populated, indicating a strong foundation for bibliometric analysis Table 1.
| Field | Missing Values |
|---|---|
| PMID | 0 |
| Title | 46 |
| Abstract | 6912 |
| Authors | 242 |
| Journal | 124 |
| PubDate | 0 |
| DOI | 2021 |
| PublicationType | 0 |
| MeSHTerms | 31902 |
| Affiliation | 2365 |
| Country | 10 |
| Year | 0 |
3.2. Temporal Trends in AI Publications
The volume of AI-related biomedical research has grown dramatically over the past decades. Before 2000, AI in this domain remained marginal, with fewer than 800 total articles published across four decades. Between 2000 and 2009, a modest uptick occurred, yielding 595 publications. From 2010 to 2014, growth continued steadily, with 424 papers published, reflecting the early integration of AI into biomedicine (Fig. 1).
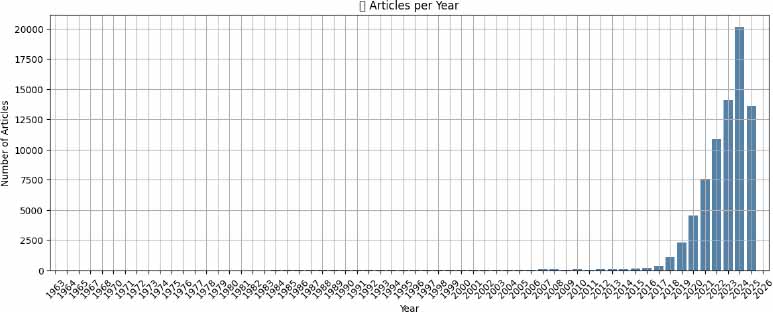
However, the period following 2015 marked a paradigm shift. Between 2015 and 2019, output surged to 4,136 articles, and in 2020 alone, 4,550 AI-related studies were indexed. The trajectory steepened further: 7,577 articles were published in 2021, 10,839 in 2022, 14,076 in 2023, and 20,135in 2024-the highest annual total on record. By the partial year of 2025, 13,617 studies had already been logged. This explosive growth likely corresponds with advancements in deep learning, increased computational resources, and heightened interest following the COVID-19 pandemic.
3.3. Geographic Distribution
According to the place of publication, the global profile of AI-driven biomedical research shows dominance by high-income nations, though emerging economies are steadily increasing their contributions. The United States leads unequivocally, contributing 24,958 articles-more than one-third of all indexed studies. This reflects both a rich research ecosystem and deep investment in digital health. England ranks second with 17,280 publications, supported by the UK’s emphasis on AI in public healthcare transformation (Fig. 2).
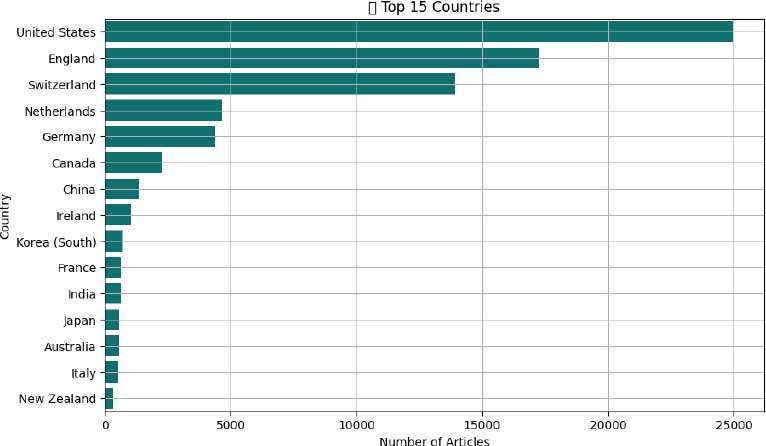
Switzerland’s output (n = 13,936) is particularly notable given its smaller population, likely reflecting concentrated innovation hubs and global pharmaceutical leadership. Other major contributors include the Netherlands (n = 4,657), Germany (n = 4,397), and Canada (n = 2,264), each backed by established biomedical and AI research infrastructures. Ireland (n = 1,045), France (n=629), and Australia (n =568) also demonstrate strong engagement. China (n = 1,355) was the 6th globally but number one in Asia. In the rest of Asia, Korea (n = 709), India (n = 625), and Japan (n = 581), reflect growing capacity, aligning with national strategies in AI and healthcare digitization. This distribution underscores both a global concentration of research in wealthier nations and a growing inclusivity as more countries invest in AI for health (Fig. 3).
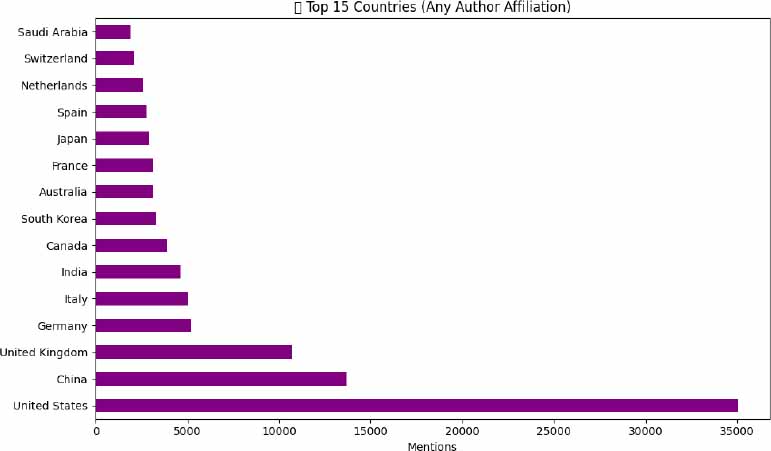
When considering the author affiliations, the landscape of global contributions to AI-driven biomedical research shifts slightly, highlighting deeper international collaboration. The United States remains the most prominent contributor, with approximately 35,000 mentions, reinforcing its central role in AI and health innovation ecosystems. China follows closely, reflecting its increasing integration into global scientific networks. The United Kingdom ranks third, driven by strategic investments in AI for healthcare delivery and academia-industry partnerships. Other significant contributors include Germany, Italy, and India, while countries such as Saudi Arabia, Switzerland, and the Netherlands also appear in the top 15, emphasizing the expanding diversity of participating nations. This distribution suggests a growing globalization of AI in medicine, with research efforts no longer concentrated solely in a few traditional scientific powerhouses.
3.4. Leading Journals
AI-focused biomedical research is distributed across a wide spectrum of journals, signaling the field’s interdisciplinary nature. Sensors (Basel, Switzerland) tops the list with 1,919 publications, indicating a strong representation of studies involving health-related sensing technologies and diagnostic tools. Scientific Reports (n = 1,832), Diagnostics (Basel, Switzerland) (n=1096), followed by Cureus (n=990) (Fig. 4).
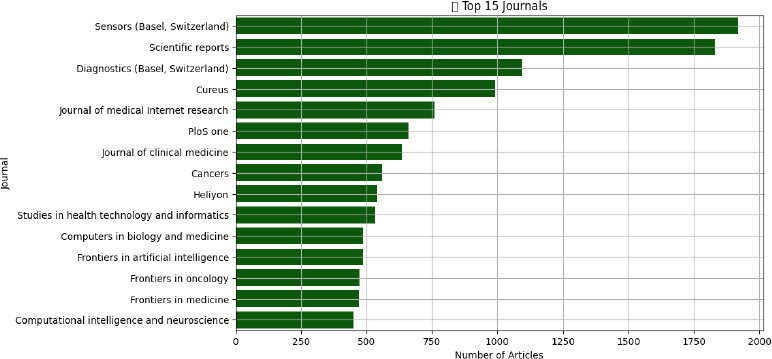
Journals like Diagnostics (n = 1096) and Cureus (n = 990) emphasize clinical translational work. Computational-focused publications such as Frontiers in Artificial Intelligence (n = 487488), Computers in Biology and Medicine (n = 947), and Computational Intelligence and Neuroscience (n = 450) provide dedicated spaces for technical innovation. Specialty journals-Frontiers in Oncology (n = 474), and Journal of Clinical Medicine (n = 635)-demonstrate the application of AI within domain-specific contexts. Journal of Medical Internet Research (n = 760) reveals the role of AI in informatics, behavior, and mental health. These findings confirm that AI research is not siloed in computational science but has penetrated core clinical and translational journals.
3.5. Thematic Trends via MeSH Terms
When focusing specifically on AI-related Medical Subject Headings (MeSH) and keywords-such as machine learning, deep learning, and neural networks-the analysis reveals the core technological themes shaping contemporary biomedical research. Machine learning stands out as the most frequently cited concept, appearing in 16,281 articles, underscoring its foundational role in predictive modeling, diagnostics, and clinical decision support systems. Deep learning follows with 12,297 mentions, reflecting widespread adoption of complex neural architectures, particularly in imaging, genomics, and signal interpretation. Neural networks, a broader term encompassing earlier architectures, remain prevalent (n = 7,117), particularly in foundational studies. Meanwhile, natural language processing (n = 1,681) continues to gain traction, driven by applications in electronic health record analysis and clinical documentation. Other rising domains include radiomics (n = 1,440), central to precision oncology, and emerging trends like transformers, GPT, Large Language Model (LLMs), and AI ethics, albeit with lower frequencies-signaling a new wave of interest in generative models and ethical considerations (Fig. 5).
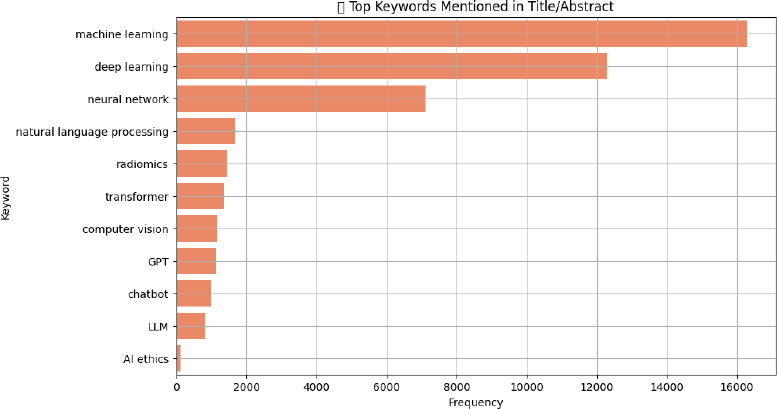
Terms like “transformers” (n = 1,358) and “GPT” (n = 1,641) signaled a strong pivot toward large language models (LLMs) in recent years. Computer vision (n = 1,137) captured AI applications in diagnostics and surgery. Chatbots (n = 1,008) and general LLMs (n = 832) illustrated the expanding presence of AI in patient engagement and digital triage. Notably, ethical concerns were comparatively scarce, with only 110 articles indexed under AI ethics. This limited presence suggests that while technical advancements are accelerating, scholarly engagement with regulatory, equity, and safety considerations remains disproportionately low.
4. DISCUSSION
Spanning 76722 publications across a 74-year timeframe (1950–2024), this bibliometric analysis provides a sweeping overview of artificial intelligence’s (AI) evolving role within biomedical research. What was once a fringe academic endeavor has now become a structural element in healthcare delivery, decision-making, and innovation. Observed not through a single metric but through intersecting trends-publication timelines, national research contributions, and journal thematic alignments-the trajectory of computational methods in biomedical domains reflects a clear shift from peripheral exploration to foundational integration [1, 7]. Interestingly, the field’s early presence prior to the year 2000 was modest, often confined to conceptual frameworks rather than applied science, with minimal clinical anchoring. Notably, the period from 2000 to 2014 didn’t explode with output, but instead built quiet momentum as machine learning tools became more refined and medical informatics matured in parallel.
What followed, however, was not simply a continuation but a transformation. By 2015, new catalysts-deep learning architectures, scalable data repositories, and cloud infrastructure-accelerated this evolution. Then came the pandemic. In the midst of COVID-19, an urgent demand for digital triage, epidemiological modeling, rapid diagnostics, and vaccine pipeline optimization brought these techniques into operational reality. That nearly 20,000 studies appeared in 2024 alone is less about trendlines and more about systems convergence-research, policy, and practice aligning around algorithmic health tools [1-4].
Spatially, the landscape is far from balanced. The most prolific outputs remain tethered to resource-rich nations-namely the United States, the United Kingdom, and Switzerland-where innovation ecosystems are supported by national investment strategies and translational research pipelines. Switzerland, despite its smaller population, exhibits remarkable productivity, likely attributable to its concentration of pharmaceutical Research and Development (R&D) hubs and collaborative biomedical clusters. Meanwhile, the rise of contributors from Asia-particularly China, Korea, India, and Japan-suggests an eastward expansion of scholarly presence. Yet, the disparity persists. The research visibility of lower-income settings remains constrained, shaped not only by access to funding but also by indexing policies, infrastructure inequality, and global publishing hierarchies that remain to be dismantled [8-11].
The spread of AI-related publications across diverse journals illustrates the field’s growing clinical relevance. Journals like Sensors (Basel, Switzerland), Scientific Reports, Diagnostics (Basel, Switzerland), and Cureus dominate the publishing landscape, indicating that AI has moved well beyond specialist technical forums. The presence of domain-specific journals in oncology, radiology, psychology, and public health further highlights the integration of AI into specialty practice and patient-facing care. This interdisciplinary diffusion signals that AI is becoming a core enabler across nearly every domain of modern medicine-from predictive oncology to behavioral diagnostics and population health [7-9].
Machine learning and deep learning continue to form the methodological core of biomedical AI, as reflected in the most frequently cited MeSH terms and keywords. A broad spectrum of clinical domains now reflects the widespread integration of machine learning and deep learning frameworks. Image analysis, digital histopathology, genomic profiling, and the construction of predictive models for patient outcomes are just a few of the many areas being influenced. At the same time, a notable evolution is unfolding in natural language processing (NLP). Transformer-based architectures-including those underpinning large language models like Generative Pre-trained Transformers (GPTs)-have carved out a new frontier in biomedical AI. These models, increasingly adopted, are not just parsing clinical narratives but also reshaping documentation workflows, automating the generation of discharge summaries, and enabling more conversational human-computer interfaces in care delivery [2, 10].
Computer vision, particularly in radiomics, plays a pivotal role in refining diagnostic imaging workflows. Meanwhile, the growth of chatbot applications signals an emerging shift towards interactive AI systems designed to improve patient engagement and streamline administrative communication. This dual trajectory-of visual data interpretation and textual automation-underscores the layered complexity of AI integration in healthcare. Far from being a uniform phenomenon, AI’s presence now spans diagnostic assistance, documentation efficiency, and real-time patient interaction [4, 11].
Yet, what remains striking is the minimal attention paid to ethical frameworks. Only 110 articles mentioned AI ethics, underscoring a dangerous lag between technological progress and ethical preparedness. This gap must be addressed urgently through both scholarly engagement and policy intervention [1-3].
While the dataset provides a rich foundation, it also reveals important structural limitations. Nearly 40% of the records lacked MeSH indexing-an omission that weakens thematic classification and complicates structured searches. A limitation primarily affecting recently published articles due to annotation lag, and non-indexed publication types such as commentaries or editorials. About 9% of records had missing abstracts, a pattern consistent with older entries (pre-1990s), conference proceedings, or errata, which are often excluded from full indexing and may be considered for exclusion in natural language processing applications. Many older entries also lacked DOI identifiers.
These metadata inconsistencies pose real barriers to automated synthesis, systematic reviews, and AI-driven literature mining. Improving consistency in indexing, especially for emerging journals and preprints, is essential for building a future-ready biomedical knowledge ecosystem [5-12]. Future efforts may further enrich this dataset by integrating cross-database identifiers (e.g., DOIs, Scopus IDs) or by linking with external metadata sources (e.g., Web of Science, CrossRef, or Dimensions.ai) [13, 14].
In addition to plotting growth patterns, this research makes unique contributions to existing bibliometric research. Compared with previous analyses, which placed main emphasis on publication volume, we assessed metadata completeness for abstracts, DOIs, and MeSH terms, identifying systematic gaps that directly affect the reliability of future bibliometric syntheses. We also incorporate thematic mapping of state-of-the-art AI methodologies, e.g., machine learning, deep learning, natural language processing, radiomics, and the recent emergence of large language models, thus reflecting the technological history of AI in biomedical science with a more granular perspective. Significantly, by emphasizing the limited focus on AI ethics and the continued lack of global representation, this analysis goes beyond descriptive statistics and offers practical advice on how to allocate funding, regulate AI, and engage in equitable health innovation based on artificial intelligence.
5. LIMITATIONS
There are a number of limitations that this research has. First, PubMed does not represent the entire range of biomedical literature, even though it offers a very extensive source of relevant literature, especially for those published only in Scopus, Web of Science, or local databases. In that case, such a dataset could be inadequate to represent the output of research in some countries and journals. Second, even when metadata was made standard and validated, inconsistencies were still present, including the most notable absent MeSH terms, abstracts, and DOIs, which could have adversely affected thematic classification and the accuracy of bibliometric mapping. Third, we might have missed studies in which the concept of artificial intelligence appeared but was not directly called by Title/Abstract search terms, despite adding keywords like machine learning, deep learning, natural language processing, and radiomics. Fourth, bibliometric analysis is descriptive in nature; thus, it can point out trends, gaps, and distributions, but cannot determine the quality, impact, or clinical significance of individual studies. Lastly, as AI research is fast-paced and dynamic, the trends presented here are not a time capsule, because publication trends can change quickly as technology advances and global health needs evolve.
5.1. Implications and Future Directions
The insights from this analysis resonate across several domains of health and science policy. For researchers, the findings clarify where innovation is happening, which journals are most receptive, and what thematic areas are surging. This can help shape publication and collaboration strategies. For institutions and funders, the data spotlight not only strengths but also blind spots-regions and topics that warrant greater investment or capacity-building. Clinicians and educators can draw on these insights to integrate AI literacy into medical training, support clinical trials using AI tools, and promote evidence-based AI adoption. Policymakers, perhaps most crucially, must heed the signal that ethical AI is underdeveloped in the literature, and global participation remains uneven. Investments in regulation, governance frameworks, and equitable infrastructure are now as essential as algorithmic refinement.
Looking ahead, the field would benefit from deeper bibliometric efforts, including longitudinal topic modeling to map conceptual evolution, network analysis of co-authorships to chart scientific collaboration, and tracking how AI regulation impacts publication trends and clinical implementation. AI in medicine is no longer emerging. It is already here. Ongoing bibliometric monitoring will be essential to ensure that its trajectory is not only innovative but also inclusive, ethical, and aligned with real-world healthcare needs.
CONCLUSION
This bibliometric analysis of 76,722 AI-related publications indexed in PubMed from 1950 to 2024 offers a comprehensive lens into the field’s rapid evolution and integration within biomedical research. The data reveal a significant inflection point beginning in 2015, driven by advances in deep learning, the emergence of large language models, and the digitization of healthcare data, resulting in exponential growth. This surge, further amplified by the COVID-19 pandemic, reflects both technological breakthroughs and mounting clinical demand for intelligent, scalable solutions in diagnostics, decision-making, and care delivery.
While the United States, England, and Switzerland continue to lead AI-driven biomedical research, growing contributions from countries like China, Japan, and India indicate a widening global footprint. The spread of AI publications across clinical, computational, and interdisciplinary journals reinforces the technology’s mainstream acceptance. Thematic dominance of machine learning, deep learning, and radiomics highlights areas of maturity, while the limited focus on AI ethics exposes a critical blind spot. As AI becomes further embedded in healthcare, ongoing bibliometric monitoring will be key to informing funding, governance, education, and collaborative innovation to ensure equitable and responsible integration.
AUTHORS’ CONTRIBUTIONS
The authors confirm their contribution to the paper as follows:. K.A. and O.A.: Study conception and design were carried out; Analysis and interpretation of results were conducted; K.A.: Data collection was performed; O.A.: Draft manuscript preparation was done. All authors reviewed the results, contributed to the discussion, and approved the final version of the manuscript.
LIST OF ABBREVIATIONS
| AI | = Artificial Intelligence |
| SQCCCRC | = Sultan Qaboos Comprehensive Cancer Care and Research Centre |
| NLM | = National Library of Medicine |
| NCBI | = National Center for Biotechnology Information |
| PMID | = PubMed Identifier |
| MeSH | = Medical Subject Headings |
| DOI | = Digital Object Identifier |
| CSV | = Comma-Separated Values |
| API | = Application Programming Interface |
| NLP | = Natural Language Processing |
| R&D | = Research and Development |
| LLM(s) | = Large Language Model(s) |
| GPT | = Generative Pretrained Transformer |
| PDAT | = Publication Date |
| PLOS ONE | = Public Library of Science ONE |
| MEDLINE | = Medical Literature Analysis and Retrieval System Online |
| LID | = Locator Identifier |
CONSENT FOR PUBLICATION
No individual-level data, personal identifiers, biological samples, images, audio, or video materials were used in this study.
AVAILABILITY OF DATA AND MATERIALS
All the data and supporting information are provided within the article.
ACKNOWLEDGEMENTS
The authors would like to thank the Sultan Qaboos Comprehensive Cancer Care and Research Centre (SQCCCRC), University Medical City, Muscat, Oman, for providing institutional support in facilitating this research.


