Data Mining Approach to Identify Disease Cohorts from Primary Care Electronic Medical Records: A Case of Diabetes Mellitus
Abstract
Background:
Identification of diseased patients from primary care based electronic medical records (EMRs) has methodological challenges that may impact epidemiologic inferences.
Objective:
To compare deterministic clinically guided selection algorithms with probabilistic machine learning (ML) methodologies for their ability to identify patients with type 2 diabetes mellitus (T2DM) from large population based EMRs from nationally representative primary care database.
Methods:
Four cohorts of patients with T2DM were defined by deterministic approach based on disease codes. The database was mined for a set of best predictors of T2DM and the performance of six ML algorithms were compared based on cross-validated true positive rate, true negative rate, and area under receiver operating characteristic curve.
Results:
In the database of 11,018,025 research suitable individuals, 379 657 (3.4%) were coded to have T2DM. Logistic Regression classifier was selected as best ML algorithm and resulted in a cohort of 383,330 patients with potential T2DM. Eighty-three percent (83%) of this cohort had a T2DM code, and 16% of the patients with T2DM code were not included in this ML cohort. Of those in the ML cohort without disease code, 52% had at least one measure of elevated glucose level and 22% had received at least one prescription for antidiabetic medication.
Conclusion:
Deterministic cohort selection based on disease coding potentially introduces significant mis-classification problem. ML techniques allow testing for potential disease predictors, and under meaningful data input, are able to identify diseased cohorts in a holistic way.
1. INTRODUCTION
Recent advances in the design and implementation of large patient-level electronic medical records (EMRs) from national primary care databases have created opportunities in clinical, epidemiological and public health research [1, 2]. In a typical primary or ambulatory care setting, large volumes of data are generated as patients go through various phases of treatment. Individual patients’ longitudinal data on demographics, lifestyle, disease and treatment history, clinical and laboratory parameters, hospitalization statistics, and clinical events are typically organized and stored in a form of relational database. Such databases present unique challenges in terms of efficient and effective extraction of data for various investigative interests [3]. One of the challenging aspects in this context is the identification of disease cohorts for retrospective or prospective clinical epidemiological studies [4, 5].
Diagnostic codes, such as the International Classification of Diseases (ICD) codes or Read codes [6], are generally used to identify disease cohorts from EMRs [4]. The reliability of diagnosis coding for various diseases has been extensively examined for many primary care databases including The Health Improvement Network (THIN) database from the United Kingdom [7-9]. However, there are four specific issues in relation to identifying cohorts by diagnostic codes: (1) differentiating between disease subtypes from high-level codes, (2) overlapping codes of disease subtypes longitudinally at individual patient level, (3) absence of codes for diseased patients (false negatives), and (4) presence of disease specific codes for patients without the specific disease (false positives).
With regards to diabetes mellitus (DM), identification and appropriate classification of different types of diabetes in the primary care databases are particularly challenging [5, 10-13]. These challenges border mostly on inaccurate coding leading to misclassification, misdiagnosis, and undiagnosed diabetes [12]. Algorithms based on laboratory, clinical, and medication data have thus been proposed as tools for distinguishing between type 1 diabetes mellitus (T1DM) and type 2 diabetes mellitus (T2DM) [10, 14-16]. However, the overall accuracy and reliability of derived disease cohorts based on diagnostic codes can be improved by implementing advanced machine learning (ML) or statistical data mining techniques and clinically guided cohort selection algorithms that robustly capture comprehensive patient level information available in the EMRs [4, 5, 12, 13].
Shivade and colleagues (2014) have conducted a systematic review of various techniques used for the identification of different disease cohorts from different sources of clinical databases [2]. Some of these proposed algorithms have been criticized for their appropriateness in the context of other studies [17]. While several studies compared or applied ML techniques to identify T2DM patients, to the best of our knowledge, there is no study that employed an extensive assessment of diagnostic codes, deterministic clinical selection algorithms, and ML algorithms simultaneously to identify T2DM cohorts from primary care EMRs.
The aims of this exploratory methodological study were to (1) explore technical challenges in the extraction of disease cohorts, (2) compare the ability of different clinically guided cohort selection algorithms to identify the disease cohorts, and (3) compare the disease cohorts identified by ML algorithms and clinically guided cohort selection algorithms using a large nationally representative primary care database from the UK.
2. MATERIALS AND METHODS
In this section, we introduce the primary care database, describe the challenges in identifying cohort of patients with specific disease (i.e. T2DM), explain the clinically guided cohort selection algorithms, and the data mining and computational processes leading to comparison of different supervised ML techniques.
2.1. Data Source
Data from The Health Improvement Network (THIN), which is a patient level primary care data from UK was used in this study. THIN is an ongoing primary care database of medical records of anonymized patients from general practitioners, covers over 600 UK general practices, and has been linked to the hospital episode statistics (HES) and other statistics from the National of Bureau of Statistics. Longitudinal patient level records have been collected since 1990 and the current version of the database holds more than 13 million individual patient records. The patients included in this database are representative of the UK population by age, gender, medical conditions and death rates adjusted for demographics and social deprivation. The accuracy and completeness of THIN database have been previously described elsewhere [18, 19]. The THIN database is considered as one of the most comprehensive patient level databases available globally, and has been extensively used by researchers and government bodies for clinical, epidemiological and public health related studies [20]. The database contains extensive information on individuals’ demographic, clinical, laboratory, medications and event history data. The study protocol was approved by the Independent Scientific Review Committee for the THIN database (Protocol Number: 15THIN030) and the Institutional Review Board of QIMR Berghofer Medical Research Institute.
2.2. Challenges in Identifying Disease Cohort
THIN uses the UK’s standard Read code classification system which is useful for hierarchical classification of patients’ specific circumstances and lifestyles, thereby enhancing scalability and retrieval (6). However, the Read coding system is complex as a disease or an encounter with a general practitioner can be coded in several ways including use of existing codes or by creating new user-defined codes [21]. In this way, considerable variation and inconsistency is introduced into the coding system as observed in the case of DM [11, 14, 22].
2.2.1. Differentiating Between Disease Subtypes
Typically, many diabetes related codes are available for a single patient, some of which are high- level codes (e.g. C10 - “Diabetes mellitus”) or disease related codes that are unspecific in the description of the diabetes type (e.g. C106.12-“Diabetes mellitus with neuropathy”). Common practice has been to exclude any high level codes [14, 23] which may lead to underestimation of the disease cohort. When it is impossible to identify disease subtype (type 1 or type 2 diabetes) from the diagnostic codes, data on surrogate markers (like glutamic acid carboxylase) could be useful, but such information is not available in THIN database. Nevertheless, combinations of available biomarkers (such as age, weight or HbA1c) and medication prescriptions have been used to distinguish types of diabetes in some studies [10, 14].
2.2.2. Longitudinally Overlapping Disease Subtypes
Patients may have different disease subtypes coded longitudinally as a result of data entry errors or biological progression of the disease. While the former can lead to any combinations of subtypes, the latter may result in developing T1DM from T2DM or T2DM from gestational diabetes. To distinguish between contradictory codes, longitudinal exploratory techniques were applied in some studies [5]. Also, the techniques described above that deal with unspecific codes may be considered. To address the issue of contradictory diagnostic codes longitudinally, the following was adopted to distinguish between T1DM and T2DM.
- Use of Read codes that uniquely distinguish between T1DM and T2DM.
- In patients with unspecific codes, or longitudinally overlapping subtypes, the following is used:
- If oral antidiabetic drug (ADD) is taken ≥ 2 months, then T2DM.
- Otherwise, if age at first available diagnosis date ≤ 18 years and insulin initiated within 1 year, then T1DM.
- Otherwise, if age at first available diagnosis date > 18 years and insulin initiated within 3 months then T1DM.
- Else T2DM.
- Patients with codes for gestational diabetes and other forms of diabetes were not include in this study
2.2.3. Absence of Codes for Patients with Disease and Presence of Codes for Patients without Disease
Data entry errors such as omissions, typing, communicating errors and patients’ temporary loss of follow-up in EMRs usually result in relatively small amount of false positive, and larger numbers of false negative patients identified by diagnostic codes. Earlier studies have addressed this complex issue by employing deterministic or probabilistic algorithms [2, 15, 16]. We further focus on this challenging aspect by comparing deterministic (clinically guided) and probabilistic (ML) cohort identification approaches.
2.3. Clinically Guided Cohort Selection Algorithms
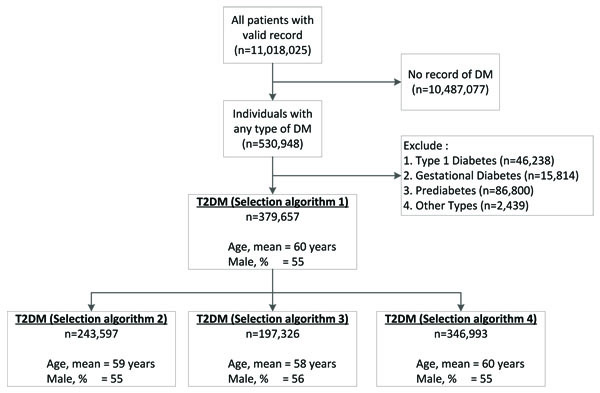
Four separate cohorts were created by applying logical, clinically guided algorithms that select patients from those who have at least one record of Read code for T2DM (Fig. 1). Specifically, the T2DM cohorts were selected on the basis of available records for:
- Selection algorithm 1: T2DM Read code (Cohort 1);
- Selection algorithm 2: Lifestyle modification intervention + T2DM Read code (Cohort 2);
- Selection algorithm 3: At least one prescription for antidiabetic medication + lifestyle modification intervention + T2DM Read code (Cohort 3);
- Selection algorithm 4: At least one prescription for antidiabetic medication or lifestyle modification intervention + T2DM Read code (Cohort 4).
Selection algorithm 1: T2DM Read code only; Selection algorithm 2: T2DM Read code + lifestyle modification advice. Selection algorithm 3: T2DM Read code + antidiabetic medication + lifestyle modification advice. Selection algorithm 4: T2DM Read code + (antidiabetic medication or lifestyle modification advice)
2.4. Supervised Machine Learning Techniques
The process of selecting one most appropriate probabilistic algorithm to identify patients with T2DM is described below.
2.4.1. Feature Selection
THIN database was mined to detect the most frequent medications, comorbidities, laboratory and anthropometric measurements among patients with T2DM identified on the basis of Read codes. The resulting 280 variables were combined with current clinical considerations, practices and guidelines for T2DM management [24], and 11 potential disease predictors were obtained through iterative process (Table 1). Correlation based Feature Selection (CFS) algorithm was applied to determine best of these predictors [25, 26]. This scheme independent attribute subset selection approach is particularly useful when attributes are correlated with one another, and with the class attribute. Bi-directional, forward and backward greedy search methods were applied using 10-fold cross-validation [27] and they all agreed on the same seven features described in Table 1.
2.4.2. Training Dataset
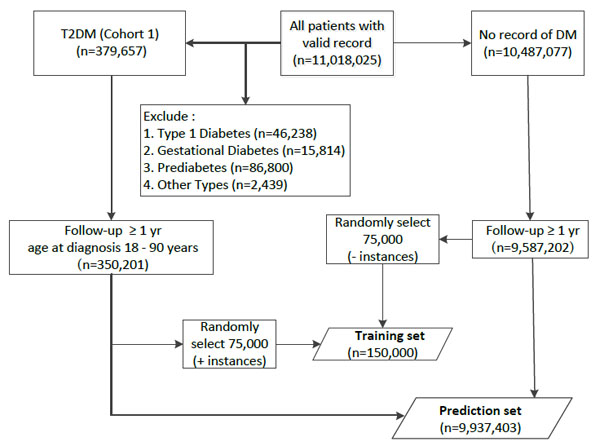
From the 11,018,025 patients in THIN database, a training dataset of 150,000 instances, containing equal number of positive and negative representatives was extracted. Positive instances were randomly selected from patients with (1) available T2DM Read code, (2) at least one year of follow-up, and (3) 18-90 years old at the time of T2DM diagnosis. Negative instances were also randomly selected from those without Read code for any subtype of DM and at least one year of follow-up (Fig. (2), training set).
| – | Feature Name | Feature Type | Selected for ML |
|---|---|---|---|
| 1 | Two measurements of HbA1c>6% or fasting blood glucose > 7 mmol/l or random blood glucose > 11.1 mmol/l within 1 year. | Binary | Yes |
| 2 | Any antidiabetic drug prescriptions for at least 6 months. | Binary | Yes |
| 3 | Average BMI. | Continuous | Yes |
| 4 | Hypertension diagnosis or antihypertensive drug use greater or equal to 6 months or beta blockers prescription for 6 months or more. | Binary | Yes |
| 5 | Chronic kidney diagnosis. | Binary | Yes |
| 6 | Retinopathy or neuropathy diagnosis. | Binary | Yes |
| 7 | Average systolic blood pressure. | Continuous | Yes |
| 8 | Lifestyle modification advice. | Binary | No |
| 9 | Average HbA1c. | Continuous | No |
| 10 | Average random glucose | Continuous | No |
| 11 | Heart failure or myocardial infarction or stroke or coronary artery disease | Binary | No |
2.4.3. Classification Algorithm Selection
Keeping the selected subset of 7 robust predictors of T2DM, six classification algorithms were applied to the training set. Ten repeat 10-fold cross-validation was applied to calculate true positive rate (sensitivity), true negative rate (specificity), and area under receiver operating characteristic curve (AUC). Percent of correctly classified instances and required central processing unit (CPU) time for training the algorithms were also derived. The algorithms for comparison were: Naïve Bayes [28, 29], Logistic regression [30], Support Vector Machine (SVM) [31, 32], Multilayer Perceptron (MP) [33], Decision Tree with J48 modification [34], and One Rule [35].
One Rule algorithm performed significantly worse. Except differences in CPU time, performance of other algorithms was similar. Among them, Naïve Bayes had lower sensitivity misclassifying approximately 500 additional patients compared to other approaches. AUC was smaller for SVM and J48, while SVM and MP required significantly higher CPU time (Table 2). Interestingly, neither body mass index nor blood pressure contributed significantly to any model. Logistic regression was selected as most appropriate model for predicting T2DM. The model obtained from full training dataset was applied to all THIN database patients with no record of Read code for diabetes diagnosis other than T2DM, and with available follow-up for at least one year (Fig. (2), prediction set).
| – | Naïve Bayes | Logistic Regression |
Multilayer Perceptron |
Support Vector Machine |
J48 Decision Tree |
One Rule |
|---|---|---|---|---|---|---|
| Percent correct | 95.6 | 95.9 | 95.9 | 95.9 | 95.9 | 91.7 |
| TPR | 0.98 | 0.99 | 0.99 | 0.99 | 0.99 | 0.99 |
| TNR | 0.93 | 0.93 | 0.93 | 0.93 | 0.93 | 0.84 |
| AUC | 0.98 | 0.98 | 0.98 | 0.96 | 0.96 | 0.92 |
| CPU time | 0.09 | 3.36 | 68.03 | 191.9 | 1.78 | 0.21 |
3. RESULTS
The distributions of basic characteristics of patients identified by all four clinically guided algorithms and the ML algorithm were similar (Table 3). Clinically guided algorithms 1-4 and the ML algorithm resulted in cohorts of 379,657; 243,597; 197,326; 346,993; and 383,330 patients with T2DM respectively. For patients identified by the ML algorithm who did not have a Read code, the first available date of entry of the significant predictors was used as their date of diagnosis. At the time of diabetes diagnosis, identified patients were on average 60 years old, 86 kg in weight with 55% male. The proportions of those who had two elevated glucose level measurements within one year were 75, 86, 90, 79, and 82% in cohorts identified by selection algorithms 1-4 and ML respectively. With median 11 years of follow-up post diagnosis, proportions of those who received at least one prescription for antidiabetic medication were 79, 81, 100, 87, and 75% in cohorts identified by rules 1-4 and ML respectively.
Among the cohort of T2DM patients identified by ML algorithm, 317,979 (83% of 383,330) patients had Read code for T2DM (Table 4). It is worth noting that 59,678 (16% of 379,657) patients with a record of T2DM Read code were not selected by ML approach. Almost a fifth (17% of 383,330) of the patients in ML cohort were without a record of T2DM Read code. Of them, 52% had at least one measure of elevated glucose level and 22% had received at least one prescription for antidiabetic medication (Table 4).
In order to assess the proportion of patients that remain undetected by the algorithms used in this study, complement cohort-specific analysis was performed (data not shown). Among patients not selected by ML as T2DM, only 884 patients had at least two elevated glucose measurements (HbA1C > 6% or fasting blood glucose > 7 mmol/l or random blood glucose > 11.1 mmol/l) within 1 year, compared to 32,039, 106,671, 137,796, and 42,583 patients not selected by selection algorithms 1-4.
| – | Selection Algorithm 1 | Selection Algorithm 2 | Selection Algorithm 3 | Selection Algorithm 4 | ML |
|---|---|---|---|---|---|
| Patients, n | 379,657 | 243,597 | 197,326 | 346,993 | 383,330 |
| Age at diagnosis (years) α | 60 (15) | 59 (14) | 58 (14) | 60 (15) | 59 (15) |
| Age at diagnosis (years) * | 61 (50,71) | 60 (50,69) | 58 (49,67) | 60 (50,70) | 60 (50,70) |
| ≤40 | 32,644 (9) | 19,761 (8) | 17,969 (9) | 29,701 (9) | 71,752 (19) |
| 41-50 | 62,656 (17) | 43,872 (18) | 39,289 (20) | 59,608 (17) | 58,813 (15) |
| 51-60 | 90,464 (24) | 62,610 (26) | 54,006 (27) | 85,587 (25) | 84,277 (22) |
| 61+ | 193,893 (51) | 117,354 (48) | 86,062 (44) | 172,097 (50) | 168,488 (44) |
| Male # | 208,155 (55) | 134,393 (55) | 110,178 (56) | 191,107(55) | 200,447 (52) |
| At least one prescription# | 300,722 (79) | 197,326 (81) | 197,326 (100) | 300,722 (87) | 287,095 (75) |
| Prescription duration ≥ 6 months# | 243,064 (64) | 171,800 (71) | 171,800 (87) | 243,064 (70) | 254,255 (66) |
| RBG (mmol/l) α § | 11.5 (5.1) | 11.4 (5.1) | 12.1 (5.3) | 11.6 (5.2) | 11.3 95.2) |
| RBG (mmol/l) α ‡ | 9.5 (3.4) | 9.4 (3.3) | 9.9 (3.4) | 9.6(3.4) | 9.1 (3.5) |
| FBG (mmol/l) α § | 8.4 (2.3) | 8.4 (2.3) | 8.9 (2.4) | 8.5 (2.3) | 8.3 (2.3) |
| FBG (mmol/l) α ‡ | 7.8 (2.1) | 7.7 (2.0) | 8.0 (2.1) | 7.8(2.1) | 7.5 (2.1) |
| HbA1c (%)α § | 8.4 (2.1) | 8.4 (2.1) | 8.7 (2.2) | 8.5 (2.2) | 8.3 (2.1) |
| HbA1c (%)α ‡ | 7.5 (1.4) | 7.5 (1.3) | 7.7 (1.3) | 7.5(1.4) | 7.4 (1.3) |
| Composite measure# ‡ | 283,419 (75) | 208,787 (86) | 177,689 (90) | 272,875 (79) | 314,574 (82) |
| Weight (kg) α § | 89.4(20.8) | 90.3 (21.0) | 91.1 (21.1) | 89.6 (20.9) | 89.3 (21.0) |
| Weight (kg) α ‡ | 85.0 (19.8) | 86.6 (19.9) | 87.6 (20.0) | 85.5 (19.8) | 86.1 (20.6) |
| BMI (kg/m2) α § | 31.6 (6.7) | 32.0 (6.7) | 32.2 (6.7) | 31.7 (6.7) | 31.7 (6.8) |
| BMI (kg/m2) α ‡ | 30.2 (6.1) | 30.7 (6.1) | 31.0 (6.2) | 30.4(6.1) | 30.7 (6.7) |
| Normal weight # | 22311(12) | 15,821 (11) | 12,339 (11) | 21,108 (12) | 24,453 (13) |
| Overweight # | 58,447 (32) | 44,283 (32) | 35,289 (31) | 55,885 (32) | 61,846 (32) |
| Grade 1 obese # | 52,465 (29) | 41,323 (30) | 33,669 (30) | 50,423 (29) | 55,684 (29) |
| Grade 2 obese # | 27,168 (15) | 22,163 (16) | 18,497 (16) | 26,336 (15) | 29,178 (15) |
| Any CVD# | 106,523 (28) | 67,011 (28) | 51,905 (26) | 96,147 (28) | 93,703 (24) |
| CKD# | 10,547 (3) | 8,035 (3) | 4,609 (2) | 9,445 (3) | 12,404 (3) |
| Cancer# | 24,159 (6) | 15,998 (7) | 11,084 (6) | 21,536 (6) | 22,112 (6) |
| Hypertension# | 149,752 (39) | 104,916 (43) | 79,193 (40) | 137,440 (40) | 140,341 (37) |
| Follow-up (years) * | 11 (6,17) | 10 (6,15) | 11 (6,16) | 11(6,17) | 10 (5,16) |
| – |
Machine Learned T2DM Cohort (n=383,330) |
|
|---|---|---|
| – | With Read Code | Without Read Code |
| Patients # | 319,979 (83) | 63,351 (17) |
| Age at diagnosis (years) α | 60 (14) | 54 (24) |
| Age at diagnosis (years) * | 60 (50, 70) | 56 (33, 73) |
| ≤ 40 | 25,645 (8) | 46,107 (73) |
| 41-50 | 56,583 (18) | 2,230 (4) |
| 51-60 | 81,262 (25) | 3,015 (5) |
| 61+ | 156,489 (49) | 11,999 (19) |
| Male # | 176,568 (55) | 23,879 (38) |
| At least one prescription # | 273,272 (85) | 13,823 (22) |
| Prescription duration ≥ 6 months # | 241,517 (76) | 12,738 (20) |
| RBG >11.1 mmol/l #, | 101,135 (32) | 1,471 (2) |
| FBG > 7 mmol/l# | 50,446 (16) | 1,695 (3) |
| HbA1c > 6%# | 274,565 (86) | 29,793 (47) |
| Composite measure# | 274,565 (86) | 29,793 (47) |
4. DISCUSSION
In this study we addressed a number of problems encountered by computer based methods in the complex tasks of identifying a disease cohort from large EMR databases. Specifically, (1) we have defined and discussed common technical challenges in differentiating diabetes subtypes, (2) combining clinical, medication and morbidity information with database patterns, we selected a set of best predictors as feeds to ML algorithms that can be used to identify patients with T2DM in the absence of any disease code, and (3) compared T2DM cohorts identified by clinically guided selection algorithm and ML algorithm. The results of this study are of particular interest to researchers who work with THIN database, however methods explored in this study are generalizable for any EMR with different disease coding systems.
Although we have seen no difference in distributions of basic characteristics among cohorts obtained by deterministic and probabilistic approaches, ML algorithms were found to be superior. With the use of selected features, we could confirm that 83% of the patients identified by the ML algorithm had a Read code for T2DM (Table 3). Those without Read code had comparable high risk as identified by the significant predictors. While 25 / 21% of patients with Read code / Read code + (medication or life style advice) for T2DM did not have at least two elevated measures of blood glucose within one year, only 18% of ML identified cohort did not have such measures. Among Read code / ML defined patients without elevated composite glucose measure, 69 / 41% did not receive ADD for at least 6 months. It is important to note that the patients without a Read code for diabetes are highly less likely to have a 2 elevated blood glucose measures within one year unless they were known to be diabetic or pre-diabetic.
Five of the six ML algorithms demonstrated similar performances in the training-testing data sets. Logistic regression approach was chosen as the best classifier for THIN database, however different feature patterns within other EMRs could potentially lead to better performance of other ML techniques to predict T2DM cohort. Tapak and colleagues [36] reported SVM as the better classifier, while Mani and colleagues [37] reported decision trees to outperform other ML algorithms. In this context it is important to mention that, ML algorithms cannot operate without meaningful data fed-in (“Garbage in, garbage out” principle). Although the use of different datasets makes it difficult for direct comparisons, a critical part of ML steps is the feature engineering or selection. Some recent studies have used large sets of variables associated with diabetes with the aim of enhancing the predictive accuracy [38, 39]. However, this may be limited by inclusion of irrelevant and redundant variables, and model overfitting in cases where number of observations are less than number of variables. While earlier studies were primarily based on clinically guided feature selection, we adopted a more holistic approach initially to identify the data driven candidates as potential predictors of T2DM from the whole database. Combining clinical knowledge and data driven candidate predictors, we ensured selection of most robust set of 7 predictors. Although selected features were not surprising, we have seen that, BMI, lifestyle modification advice and hypertension did not contribute to the models, while microvascular complications did.
We have compared the performances of six classification algorithms on a set of 150,000 instances, which was reconfirmed to be large enough by assessing the performance curves of several incremental classifiers. Nevertheless, training dataset was small compared to the whole database; therefore in order to ensure that our results are not prone to selection bias, we performed same analyses on 2 other randomly selected training datasets and obtained almost identical results.
Unlike most ML applications that focus on training to ensure best fit for future predictions, in this study, we have used various techniques to correct available labelling with ultimate goal to improve quality of diseased cohort (Type 2 Diabetes). It would be of great interest to compare ML error, Rule-based error, and human error in terms of predicting disease from available data. For this task a “gold standard” dataset would consist of random patients whose true disease state was reconfirmed approaching both clinician and patient. We were not able to conduct this task, as the THIN database contains de-identified patient-level data, which is true for all large EMR databases that are used for research purposes. THIN database also does not have data on surrogate markers that could improve quality of the cohort identification algorithms. Miscoding between type 1 and type 2 diabetes in the primary care database is not uncommon [40, 41]. It is important to mention that ML techniques may poorly distinguish between disease subtypes without incorporating additional classification rules. We have excluded patients with other diabetes Read codes from the dataset on which our ML algorithm was applied. Furthermore, for patients identified as T2DM without Read codes, the ML techniques are not able to provide exact diagnosis date, therefore requiring incorporation of additional techniques.
CONCLUSION
Careful investigation of diagnostic codes patterns within the databases is essential prior to conducting analyses on the disease cohort. Direct extraction of a disease cohort using diagnostic codes may lead to inclusion of falsely diagnosed patients and omitting patients with true disease state. Rule-based techniques represent conservative approach, which results in minimizing only false positive cases. ML techniques that minimize both false positives and false negatives cases represent more robust approach. However, ML techniques heavily rely on the meaningful input and use diagnostic codes for training purposes. Combining human expertise and machine power represent best strategy that allows to test hypotheses on potential disease predictors, lower human interventions, and to reduce the burden of selection bias.
LIST OF ABBREVIATIONS
| ADD | = Antidiabetic Drug |
| AUC | = Area Under the Curve |
| BMI | = Body Mass Index |
| CHD | = Coronary Heart Disease |
| CPU | = Central Processing Unit |
| CVD | = Cardiovascular Disease |
| DM | = Diabetes Mellitus |
| EMR | = Electronic Medical Record |
| FBG | = Fasting Blood Glucose |
| GP | = General Practitioner |
| HbA1c | = Glycated Haemoglobin |
| HES | = Hospital Episode Statistics |
| HF | = Heart Failure |
| ICD | = International Classification of Diseases |
| MI | = Myocardial Infarction |
| ML | = Machine Learning |
| MP | = Multilayer Perceptron |
| PAD | = Peripheral Artery Disease |
| RBG | = Random Blood Glucose |
| SD | = Standard Deviation |
| SVM | = Support Vector Machine |
| T1DM | = Type 1 Diabetes Mellitus |
| T2DM | = Type 2 Diabetes Mellitus |
| THIN | = The Health Improvement Network |
| TNR | = True Negative Rate |
| TPR | = True Positive Rate |
| UK | = United Kingdom |
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
The study protocol was approved by the Independent Scientific Review Committee for the THIN database (Protocol Number: 15THIN030) and the Institutional Review Board of QIMR Berghofer Medical Research Institute.
HUMAN AND ANIMAL RIGHTS
No Animals/Humans were used for studies that are base of this research.
CONSENT FOR PUBLICATION
Not applicable.
CONFLICT OF INTEREST
Sanjoy K. Paul has acted as a consultant and/or speaker for Novartis, GI Dynamics, Roche, AstraZeneca, Guangzhou Zhongyi Pharmaceutical and Amylin Pharmaceuticals LLC. He has received grants in support of investigator and investigator initiated clinical studies from Merck, Novo Nordisk, AstraZeneca, Hospira, Amylin Pharmaceuticals, Sanofi-Avensis and Pfizer. Ebenezer S. Owusu Adjah, Olga Montvida, and Julius Agbeve. have no conflict of interest to declare.
ACKNOWLEDGEMENTS
Sanjoy K. Paul conceived the idea and was responsible for the primary design of the study. Ebenezer S. Owusu Adjah , and Olga Montvida significantly contributed in the study design. Julius Agbeve conducted the primary raw data extraction. Ebenezer S. Owusu Adjah and Olga Montvida jointly conducted the data extraction, data manipulation, statistical analyses and developed the first draft of the manuscript. Ebenezer S. Owusu Adjah , Olga Montvida , Sanjoy K. Paul, and Julius Agbeve contributed to the finalization of the manuscript. Sanjoy K. Paul had full access to all the data in the study and is the guarantor, taking responsibility for the integrity of the data and the accuracy of the data analysis. Ebenezer S. Owusu Adjah was supported by QIMR Berghofer International Ph.D. Scholarship and The University of Queensland International Scholarship. Olga Montvida was supported by the Queensland University of Technology International Scholarship. No separate funding was obtained for this study. Melbourne EpiCentre gratefully acknowledges the support from the Australian Government’s National Collaborative Research Infrastructure Strategy (NCRIS) initiative through Therapeutic Innovation Australia and the research project funding from the National Health and Medical Research Council of Australia (Project Number: GNT1063477). Olga Montvida acknowledges the support from her associate supervisors Prof. Ross Young and Prof. Louise Hafner.


