Machine Learning Techniques used for the Histopathological Image Analysis of Oral Cancer-A Review
Abstract
Oral diseases are the 6th most revealed malignancy happening in head and neck regions found mainly in south Asian countries. It is the most common cancer with fourteen deaths in an hour on a yearly basis, as per the WHO oral cancer incidence in India. Due to the cost of tests, mistakes in the recognition procedure, and the enormous remaining task at hand of the cytopathologist, oral growths cannot be diagnosed promptly. This area is open to be looked into by biomedical analysts to identify it at an early stage. At present, with the advent of entire slide computerized scanners and tissue histopathology, there is a gigantic aggregation of advanced digital histopathological images, which has prompted the necessity for their analysis. A lot of computer aided analysis techniques have been developed by utilizing machine learning strategies for prediction and prognosis of cancer. In this review paper, first various steps of obtaining histopathological images, followed by the visualization and classification done by the doctors are discussed. As machine learning techniques are well known, in the second part of this review, the works done for histopathological image analysis as well as other oral datasets using these strategies for growth prognosis and anticipation are discussed. Comparing the pitfalls of machine learning and how it has overcome by deep learning mostly for image recognition tasks are also discussed subsequently. The third part of the manuscript describes how deep learning is beneficial and widely used in different cancer domains. Due to the remarkable growth of deep learning and wide applicability, it is best suited for the prognosis of oral disease. The aim of this review is to provide insight to the researchers opting to work for oral cancer by implementing deep learning and artificial neural networks.
1. INTRODUCTION
Over the past decades, a continual evolution associated with cancer analysis has been performed [1]. To find out the types and stages of cancer, scientists have developed different screening methods for early stage diagnosis [2-7]. An enormous quantity of cancer information has been gathered with the introduction of new technologies and is accessible to the medical research community [8]. But one of the most challenging tasks for the doctors is to accurately predict the type of cancer. Therefore, several machine learning techniques are used by medical researchers. These techniques are capable of discovering patterns and relationships among them and can efficiently predict the future outcomes of a cancer type from complicated datasets.
As machine learning techniques are more popular, a review of studies using these methods to predict oral cancer is presented.
Many characteristics that are connected with cell morphology can be acquired from digital histopathological images. It, thus, represents one of the systems’ fundamental steps to classify cells regardless of context [9]. Numerous supervised [10] and unsupervised [11] machine learning algorithms have been put forward in latest years for the classification of histopathological images like support vector machines [12, 13], neural networks [14], decision tree [15], fuzzy and genetic algorithms [16], k-NN [17, 18], kernel PCA [19], etc. These models can be extensively used for other areas of medical science, such as medicine and clinical research [20].
The neural network has developed a new area of science that is different from today's computer algorithmic calculation method [21]. The neural network is inspired by the biological neural structures and has a naiver framework. Many developed neural network systems follow some eminent features of the learning ability of biological neural networks [22, 23]. Instead of neural, physiological approaches, engineering approaches are incorporated for developing other features. The neural network by learning ability can produce new data and discover new outputs. The neural network observes learning samples, generalizes them, and produces a learning rule from the samples. The neural networks can use the learning rules to decide on any of the samples that were notseen before [24].
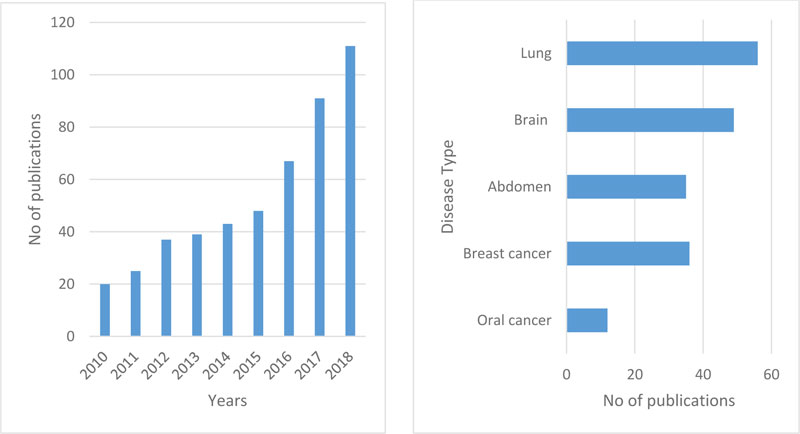
Fig. (1a) reflects the number of articles on oral cancer from the last nine years (2010-2018, including various kinds of information sets such as genomics, molecular, clinical, microarray, etc.). The use of machine learning methods in histopathological cancer images has been created relevant to a comprehensive search. The study demonstrates a fast rise in articles that have been released in the last decade based on ML apps in cancer prediction. It is still an open area to be explored. Fig. (1b) reflects paper for multiple types of cancer that have been published over the previous five years using deep learning with histopathological images. In oral cancer, only a few papers were published. Although the application of deep learning techniques is quite high for other types of cancer, the use of histopathological images of oral cancer has accomplished much less work.
For complex computer vision assignments, such as object detection, the Convolutional Neural Networks (CNN) were implemented with excellent achievements in deep learning techniques [25]. The latest studies have shown that CNNs can also be used as an efficient method for low-level image processing problems, such as restoration [26], denoising [27], [28], and mitosis detection [29]. This research discusses the classification of oral cancer as well as other cancers from the H&E color stained tissue pictures. The classification outcome can be used as the input for other assignments like the extraction of the nucleus feature classification or diagnosis.
Heterogeneous types of data like clinical, imaging, and genomics can provide enough information to identify the cancerous tissue [30-32]. The clinical examination allows for direct visualization, but a deep disease extension cannot be evaluated [33, 34]. Cross-sectional testing is becoming the basis for the pre-treatment assessment of these cancers and offers precise data on the extent and depth of the illness that can assist pathologists to determine the suitable management approach and indicate a prognosis [35].
2. ORGANIZATION OF THIS PAPER
The rest of this document is arranged as follows. An overview of oral cancer and histopathology is provided in section three. A machine learning overview is provided in the fourth section. The fifth section is the literature review that discusses various machine learning methods used for oral cancer and deep learning for other cancer kinds. Section six is the discussion that addresses the challenges and future scope. The conclusion is in the last seventh section.
3. OVERVIEW OF ORAL CANCER AND HISTOPATHOLOGY
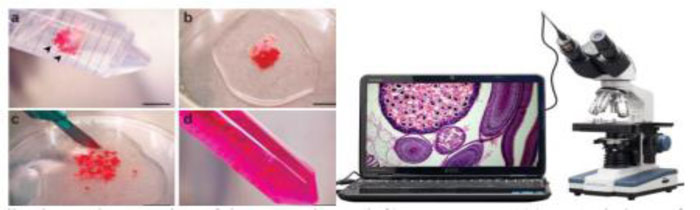
Histopathology is the human tissue analysis for a particular disease. This analysis method goes approximately as follows in clinical practice. First, a biopsy is drawn from a patient and sent to the pathology laboratory. Then a glass slide is prepared by coloring the tissue. The purpose of staining is to highlight the different structures of the tissue. For instance, staining tissue from hematoxylin and eosin (H&E) provides a dark purple color to nuclei and pink color to other structures. A pathologist can examine the tissue using a microscope after the tissue is ready and stained [36]. This total workflow is represented in Fig. (2).
Oral cancer relates to cancer occurring in the region of the head and neck [37]. India represents 86% of instances of oral cancer [38]. The most prevalent cancer in both males and females is oral cancer. The primary cause of oral cancer is chewing or smoking tobacco, a disease that claims the life of 10,000 individuals every year. Oral cancer begins in the mouth cells of the oral cavity and consists of many components such as lip, tongue, hard palate (mouth roof), mouth floor, gums, and teeth. In recent years, oral cancers have increased and further new cases are reported each year. Although the oral cavity is readily available for inspection, there is no diagnosis of oral squamous cell carcinoma (OSCC) until an advanced phase of illness has been detected [39-43]. At stage III or IV, nearly two-thirds of OSCCs are diagnosed with spread to neighboring tissues and regional lymph nodes, resulting in a poor overall survival rate of 5 years [44-47].
Oral cancer incidence is the highest in the world and the prevalence in females is smaller than in males (Global Oral Cancer Incidence Data, 2016) [48]. Oral cancers can occur in various kinds, such as squamous cell carcinoma, verrucous carcinoma, minor carcinoma of the salivary gland, lymph epithelial carcinoma, etc. About 90% of oral diseases are squamous cell carcinoma (SCC) [49]. In the ordinary situation, the throat and mouth are filled with stratified squamous cells. There is only one layer in contact with the basement membrane of this squamous cell. The epithelial layer's structural integrity is preserved by adhering to one layer. When cancer affects, this structural integrity is lost. SCC can be classified as well-differentiated SCC (WDSCC), mildly differentiated SCC (MDSCC), and badly differentiated SCC (PDSCC). SCC grading relies on how simple it is to recognize indigenous squamous epithelium features (for e.g., intercellular bridges, keratinization, etc.) [50].
A physician actually observes the suspected lesions. If he or she realizes any cancerous lesion, he indicates that the patient has to go for a confirming test, such as a biopsy. Biopsy sections prepare stained slides of hematoxylin and eosin (H&E) and are then found under the microscope. The irregularities (i.e. the various objects on the slide, including cell organization, atomic size and shape, cell size and shape, etc.) are observed by a pathologist to diagnose cancer through the microscope. An oncologist will give the patient a diagnosis depending on the pathologist's report. Fig. (3) shows the entire workflow. It is, therefore, essential for such a study to be as precise as possible. However, for each unit of the slide and feature analysis, the entire manual observation process is tedious and requires a great deal of experience. In addition, the report can also be observer biased. Because of these problems, a computerized scheme of the above-mentioned method can decrease bias and time, thereby increasing the precision of feature assessment. Therefore, the need for the hour is a computer-aided diagnostic (CAD) system for the prognosis of oral cancer, particularly in a developing and resource-poor nation such as India. It will assist in high-volume laboratories and in cancer screening camps where most instances can be benign and where the pathologist can focus mainly on the instances diagnosed by the system as malignant. Computer aided histopathological study has been conducted for various cancer detection and grading applications [51, 52].
The following Fig. (4). shows a microscopic view of the tissue sample stained with Hematoxylin and Eosin.

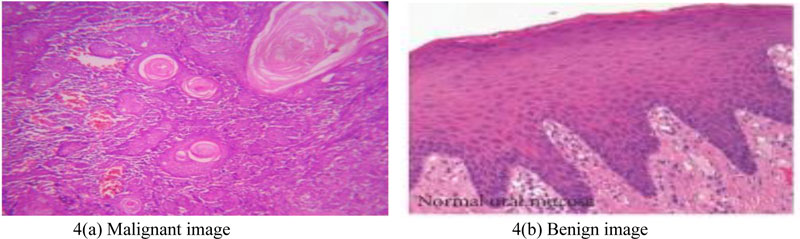
Oral mucosa has typical epithelial architecture, a usual sub-epithelial region in a healthy and precancerous tissue sample (Fig. (4b)), whereas in OSCC abnormal epithelial, keratin region and keratin-pearl structures are observed (Fig. (4a)). Invasive or non-invasive forms of tissue samples may be analyzed from an epithelial area by distinguishing the basal epithelium layer. Sub-epithelial layer features suggest precancerous stages, such as OSF. Keratin with keratin-pearl area is quantified for classification of OSCC grading.
4. OVERVIEW OF MACHINE LEARNING MODELS
Machine Learning is the ability to obtain information through Artificial Intelligence (AI) systems. The following is a proper definition of what learning requires: “a computer program is said to learn from experience E with respect to some class of tasks T and performance measure P if its performance at tasks in T, as measured by P, improves with experience E” [53]. Typically, machine learning algorithms are categorized into the following three types: supervised, unsupervised, and reinforcement learning.
(i) A dataset that includes a collection of features is provided for supervised learning algorithms. For each sample, labels or target values are also given. This mapping of features to target value labels is where the information is encoded. Once learned, it is predicted that the algorithm will determine the mapping of unseen sample features to their appropriate labels or target values.
(ii) In unsupervised learning, the aim is to obtain significant depictions and clarify the key features of the data. In this case, learning from the data does not include labels or target values.
(iii) An AI system interacts with a true or virtual environment in reinforcement learning algorithms. This provides feedback between the learning scheme and the interacting environment that is helpful in the task being learned to enhance efficiency.
The ML models i.e., Support Vector Machine (SVM), Artificial Neural Network (ANN), Decision Tree (DT), Naïve Bayes (BN), Random Forest (RF) and K-Nearest Neighbor (KNN), which are applied in this review for oral cancer prediction and diagnosis, are discussed. SVMs are widely used in the area of cancer research. Firstly, SVM maps the input vector into a higher dimensional feature space and separate the data points into two groups. This maximizes the marginal distance between the decision hyperplane and the instances nearest to the boundary. This classifier archives tremendous generalizability and can be used to accurately identify new samples. The hyperplane acts as the decision boundary between the two groups and detects any misclassification produced by the model.
In ANN, neural connections of multiple hidden layers are present. This model is considered as a gold standard method for many classification tasks, but has certain limitations. It is a time consuming technique due to the generic layered structure and considered as a “black-box”. It is difficult to detect how it is performing the classification.
DT is a tree-structured model where the input variables are represented by nodes and decision outcomes are the leaves. Depending on the structure of the DTs, they are easy to understand and widely used for many classifications tasks. When a new sample comes, the DT will be traversed for classification to speculate about its class.
BN classifiers generate conditional probability estimations. It is widely used in various tasks of knowledge discovery and reasoning due to a directed acyclic graph, which represents the probabilistic correlations amid the variables.
RF can be used for both classification and regression. It generates decision trees based on the data samples and then gets the predictions for each of them. The best solution is finally selected by the voting method. It provides better accuracy due to multiple decision trees predictions. This method works well on a large dataset with a large number of attributes.
KNN is a simple ML algorithm used for both classification and regression problems. It uses data points and identifies new data based on the measure of similarity (e.g. distance function). The classification of its neighbor is achieved by the majority vote.
Deep learning is a subset of machine learning that normally requires learning depictions at distinct levels of hierarchy to allow complex concepts to be constructed from more naive ones.
Machine learning algorithms require labeled data, complex queries involving huge amounts of data are not suitable for solving. The real application of deep learning is on a much larger scale.The following paragraphs discuss the most appropriate deep learning technologies currently available for various fields.
In the early 1980s, LeCun initially put forward the Convolutional Neural Network (CNN). It was further altered in the 1990s by LeCun and his associates [54]. CNN is a deep, feed-forward artificial neural system that is essentially connected in visual patterns. CNN does the task of machine learning in the first part, like feature extraction, and learning, and thereafter pattern grouping.
CNN is predominantly scheduled for 2-D visual pattern recognition [55]. It is widely accepted that CNN works perfectly for image problems and outperformed most of the other methods in image classification [56, 57] and segmentation tasks
Out of the different deep learning models (RNN, Autoencoder, DBN etc.), the CNN model is receiving a lot of attention in digital image processing and computer vision. There are various kinds CNN architectures, such as LeNet [11], AlexNet [56], ZFNet [58], VGGNet [59], GoogleNet [53, 60], Dense CNN [61], FractalNet [62], and Residual Network [63]. LeCun et al. proposed LeNet, which was applied for handwriting recognition. AlexNet is widely used for object detections. After that ZFNet, VGGNet, and GoogleNet were put forward based on AlexNet [64-69].
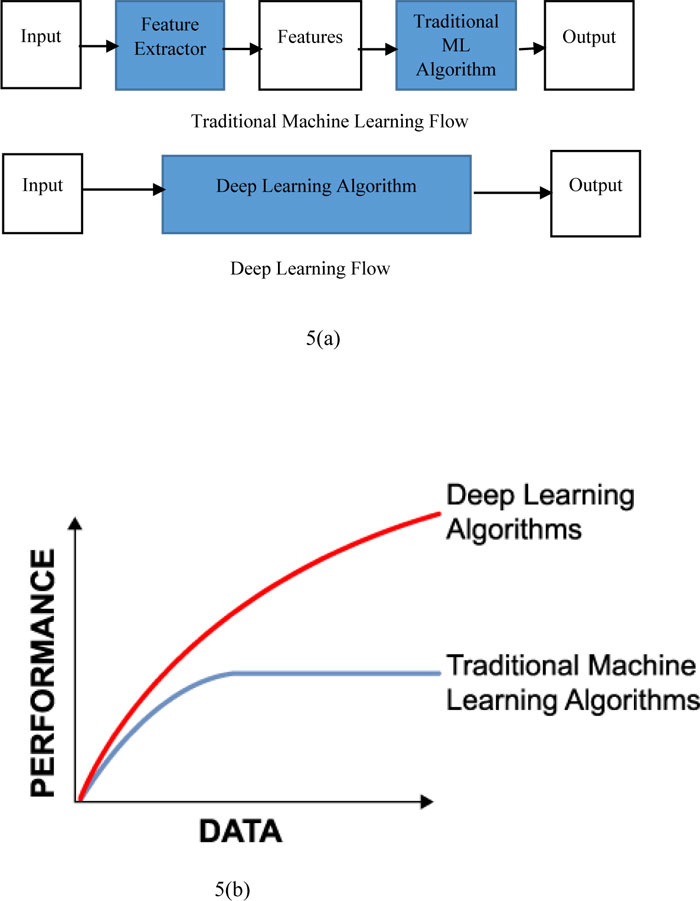
Fig. (5) shows the difference between machine learning and deep learning. It also shows that traditional machine learning is dependent on the handcrafted features whereas deep learning extracts all coarse features just like a black box. Fig. (5a) represents the performance with respect to the amount of data for deep learning as well as machine learning. Traditional machine learning is giving a better performance with a small amount of datasets. If the quantity of data rises, after a certain stage, the efficiency of traditional approaches to machine learning becomes constant. The efficiency of deep learning methods, on the other hand, improved with regard to the increase in data volume.
5. LITERATURE REVIEW
A comprehensive review of the research studies in early detection of oral cancer is discussed below in Table 1 and 2.
Rahman et al. (2017) [75] research analyzed texture based abnormalities that exist in oral squamous cell carcinoma by using histopathological slides. Histogram and Gray Level Co-occurrence Matrix (GLCM) methods were used with normal and malignant cells to extract texture features from biopsy images. They used linear SVM for automatic classification of oral cancer and obtained 100% accuracy.
Muthu et al. (2011) [72] proposed a method for “Textural characterization of histopathological images for oral sub-mucous fibrosis detection”. They used a combination of different features of wavelet family with SVM classifier that gave better accuracy.
Muthu et al. (2012b) [71] proposed “Automated oral cancer identification using histopathological images: a hybrid feature extraction paradigm”. They analyzed the texture features using HOS, LBP, and LTE and obtained an accuracy of 95.7% using the Fuzzy Classifier. Authors have shown the efficiency of Fuzzy classifier compared to DT, GMM, PNN, and KNN classifiers as the features extracted are accurate and efficient. Muthu et al. (2012a) [70] proposed a method for classification by using SVM and GMM classifier. With a very large pixel size image (1388x1040) and by applying different techniques, such as image pre-processing, segmentation, feature extraction and classification, SVM gives better results compared to other classifiers.
A study carried out by Anuradha. K et al. (2013) [35] is a method for statistical feature extraction for the detection of oral cancer. They used tumor detection by marker-controlled Wat- ershed segmentation. Energy, entropy, contrast, correlation, homogeneity are the characteristics obtained with Gray Level Co-occurrence Matrix (GLCM) through extraction. Using SVM classifiers, they acquired a precision of 92.5 percent.
Dev Kumar et al. (2015) [74] developed a model to detect the keratinization and keratin pearl areas from which the keratinization index was calculated. This Keratinization scoring index can be used as a quantitative measure for OSCC for even very low (4x) magnification. Again Dev Kumar et al. (2018) [76] proposed a method for “automatic identification of clinically relevant regions from oral tissue histological images for oral squamous cell carcinoma”. They used 12 layered deep convolution neural network (CNN) for the segmentation of different layers, such as keratin, epithelial, and subepithelial. From the segmented keratin regions, keratin pearls were detected with the texture based feature (Gabor filter) and random forest classifier, which gives 96.88% accuracy.
T.Belvin et al. (2013) [73] used backpropagation based ANN for multi-class classification of oral lesions by using run-length features and texture features. The proposed model aims for a more illustrative and patient-specific approach by using a combination of GLCM and GLRL features with an accuracy of 97.92%.
The above studies represent the recent work done for the prognosis and prediction of oral cancer using machine learning techniques. A systematic analysis is performed about the use of ML techniques for oral cancer. Depending upon the search criteria about the oral cancer prognosis, a large number of articles were obtained. A substantial rise in the number of papers published over the last ten years was noticed. Most of these studies use or combine various datasets: genomics, clinical, histopathological, biological, demographic, epidemiological data, etc. Papers based on predicting the development of oral cancer by traditional statistical approaches (e.g. chi-square, t-test) were excluded from this study. Though a full coverage of the literature cannot be achieved, the most recent and relevant publications those use ML techniques were filtered out and included in this study.
From Table 1. authors claimed that using ML techniques for the classification of oral lesion results in adequate and effective decision making. Among the different ML algorithms used for the oral cancer prediction, it was found that SVM classifier is more used due to its accurate predictive efficiency. Apart from SVM, other ML Techniques such as ANN, Decision tree, Random forest, and KNN are used. For evaluation purposes, mainly ten-fold cross validation was used after the execution of each ML process. Furthermore, accuracy, sensitivity, and specificity were also measured. The authors analyzed the ROC curve for evaluation purposes.
| Author | Images used | Features | Classifier | Performance measure | Results | Findings |
|---|---|---|---|---|---|---|
| Muthu et al. (2012a) [70] | Normal-341 OSF-429 images obtained from more than 20 patients. Image size:1388x1040 Source: School of medical science and technology, Kharagpur |
Morphological and textural features | SVM GMM |
Accuracy (99.66%) Accuracy (90.37%) |
Classifies the images into Normal, OSFWD, and OSFD | Image pre-processing, segmentation, feature extraction, and classification gives a very effective result with SVM. |
| Muthu et al. (2012b) [71] | Normal-90 OSFWD-42 OSFD-26 Image size:1388x1040 Source: “School of medical science and technology, Kharagpur” |
HOS, LBP, LTE |
Fuzzy DT GMM KNN PNN |
Accuracy (95.7%) Accuracy (77.5%) Accuracy (78.3%) Accuracy (71.7%) Accuracy (71%) |
Classifies the images into Normal, OSFWD, and OSFD | Out of different classifiers fuzzy gives promising performance because it can manage imprecise non-linear parameters when supplied with a mixture of texture and HOS function. |
| Muthu et al. (2011) [72] | Normal-90 OSFWD-42 OSFD-26 Image size:1388x1040 Source:” School of medical science and technology, Kharagpur” |
71 features of wavelet family(LBP,Gabor, BMC) | SVM | Accuracy (88.38%) | Classifies the images into Normal, OSFWD, and OSFD | A combination of different features of wavelet family gives better accuracy than when tested individually. |
| Anuradha. K. et al.2013 [35] | - | Energy, entropy, Contrast, Correlation, Homogeneity |
SVM | Accuracy (92.5%) | The classifier gives the result of the tumor as benign or malignant | SVM with GLCM features gives better results. |
| T.Belvin et al.(2013) [73] | 16 malignant images with 192 patches were collected from Himalayan Iinstitute of medical institute, Deradun | Texture features and Run Length features | Backpropagation based ANN | Accuracy (97.92%) | The combination of GLCM and GLRL features are used for the multiclass classification of oral lesions. | The proposed model aims for more illustrative and patients specific approach with ANN-based multiclass classification. |
| D.Dev kumar et al.(2015) [74] | 10 OSCC patients with 30 labelled images were collected from Tata medical centre, Kolkata | Keratinization and keratin pearls | Keratinization scoring index | Segmentation accuracy (95.08%) | Keratinization and keratin pearls were detected by keratinization index (CKI) to grade the OSCC into I,II,III. | Keratinization scoring index can be used as a quantitative measure for OSCC for even very low (4x) magnification. |
| Rahman et al. (2017) [75] | Center-1 Normal-110 Malignant-113 Center-2 Normal-86 Malignant-88 Image size(2048x1536) Source:”Ayursundra healthcare pvt Ltd and Dr. B Borooah cancer Research institute” |
94 features(homogeneity, mean, variance, energy, entropy etc. |
SVM | Accuracy (100%) | The images are classified as normal or malignant | Image size (2048x1536) and magnification of 400x gives very high performance. |
| D.Dev kumar et al.(2018) [76] | High grade-15 Low grade-25 Healthy-2 Image size:2048×1536 Total 126 images collected from “Barasat Cancer Research & Welfare Centre, Barasat, WB, India” |
Identification of various layers –epithelial,subepithelial, keratin region and keratin pearls | CNN, Gabor filter, Random forests |
Accuracy (96.88%) | Different layers of oral mucosa such as epithelial, subepithelial, and keratin and keratin pearl are identified by segmentation. | Two stage CNN for segmentation gives better prediction and outcome with valid accuracy. |
All of the studies achieved very promising results. Moreover, the selection of the most suitable algorithm depends on several parameters like types of data obtained, dataset size, time limits, and the quality of predictive performance.
A common limitation from the studies included in this review is the small size of the dataset. To model a disease classification scheme, a sufficiently large training dataset is required so that proper partitioning of training and testing sets can be made for accurate validation. Apart from the dataset size, the dataset quality and appropriate feature selection procedure are important for accurate and effective cancer prediction.
Since the team's overwhelming success with deep learning at the 2012 ImageNet Large Scale Visual Recognition Competition (ILSVRC), most image recognition methods have been substituted by deep learning (Krizhevsky et al. [12]). In Table 1, although several works on oral cancer has been proposed using machine learning techniques, very less work has been performed using deep learning for histopathological image analysis. Table 3 demonstrates the histopathological analysis of other cancer types using deep learning and Table 2 includes the research performed for oral cancer using heterogeneous data types. Even though these works do not fit in with the general statement of histopathological image analysis of oral cancer, it was decided to include them in this study due to the efficient models developed using heterogeneous data types in the field of oral cancer diagnosis. In Table 3, the wide applicability of histopathological analysis of other types of cancer is highlighted. To overcome the limitations mentioned above and the wide spread use of deep learning models for prediction in cancer domain, researchers can develop new methods by using deep learning techniques for oral cancer to achieve accurate disease outcomes. A large number of valid publicly available datasets of oral cancer can be analyzed for future cancer prediction.
Table 2 represents the different types of work done by taking heterogeneous type of data, such as clinical, genomic, molecular, etc. Some related work performed using machine learning methods for oral cancer based on this study are presented. Regarding the output, the authors claimed that the BN classifier performs better with the direct input of the integration of mixed data. BN classifier is very fast and easy to implement. SVM classifier outperforms with the combination of a clinical and molecular dataset. Complex features are learned by the kernels of SVM, however, they are very slow. Halicek M et al. (2016) and Shams W.K. et al. (2016) used the deep learning models for oral cancer prognosis with hyperspectral imaging and gene expression datasets. Remmerbach, T.W et al. used SVM classifier for the oral cancer diagnosis using matrix-assisted-laser-desorption/ ionisation-time-of-flight-mass-spectrometry mass spectrometry with 100% sensitivity and 93% specificity. Their screening method is done by a brush biopsy, which is less invasive compared to the conventional histology biopsy. Ashizawa, K et al. performed probe electrospray ionization mass spectrometry analysis of head and neck by using partial least square –logistic regression technique. The authors claimed that their method could diagnose head and neck SCC more accurately and rapidly compared to the time consuming procedure of slide preparation then microscopic assessment of pathological diagnosis. Therefore, this mass spectrometry is considered as a powerful label-free tool to analyze the molecular species, such as proteins, peptide, glycans, metabolites, etc., from a variety of biological samples. Authors have proved the efficiency of their models by comparing them with other conventional state-of-the-art models. Deep learning models have their own advantages compared to machine learning.
| Publication | Method | Cancer type | Type of data | Accuracy |
|---|---|---|---|---|
| Rosado P et al.(2013) [30] | SVM | Oral cancer | Clinical, molecular | 98% |
| Chang S-W et al.(2013) [31] | SVM | Oral cancer | Genomic, Clinical | 75% |
| Exarchos K et al.(2012) [32] | BN | Oral cancer | Imaging, Clinical, tissue genomic, blood genomic | 100% |
| Halicek M et al.(2016) [77] | CNN | Head and Neck | Hyperspectral Imaging | 96.4% |
| Shams W.K. et al.(2016) [78] | DNN | Oral cancer | Gene expression | 96% |
| Remmerbach, T.W et al.(2011) [57] | SVM | Oral cancer | Mass Spectrometry | Sensitivity=100% Specificity=93% |
| Ashizawa, K et al.(2017) [79] | Partial least squares-logistic regression | Head and Neck | Mass Spectrometry | Positive-ion mode ACC=90.48% Negative–ion mode ACC=95.35% |
| Publication | Method | Cancer type/Area | Findings |
|---|---|---|---|
| Xu et al.(2014) [80] | Multiple instance learning framework with CNN features | Colon Cancer | Patch level classification |
| Bychkov et al.(2016) [81] | Extracted CNN features from epithelial tissue | Colorectal cancer | Outcome prediction of the cancer |
| Litjens et al. (2016) [82] | fCNN based pixel classifier | Prostate and Breast cancer | Detection of Prostate and Breast cancer |
| Harsitha S.et al. (2017) [83] | AlexNet | Abdomen Cancer | Cancer classification and necrosis detection |
| Li W et al.(2018) [84] | Region based CNN | Prostate cancer | Epithelial cells detection and Gleason grading |
| Asami Y et al.(2017) [85] | Deep CNN | Brain Tumor | Grading disease stages |
| Neeraj K et al.(2017) [86] | Two separate CNN | Prostate cancer | Detection of individual nuclei and classification |
| Harsitha S.et al(2016) [87] | CNN | Abdomen cancer | Cancer classification based on immunohistochemistry and necrosis detection |
| Atsushi T.et al.(2017) [88] | Deep CNN | Lung Cancer | Accurate classification of cancer types. |
| Shidan W.et al.(2018) [89] | Deep CNN | Lung Cancer | Automated tumor recognition system |
| Teresa A.et al.(2017) [90] | CNN and SVM | Breast cancer | Four classes of cancer classification |
| Alexander R.et al.(2018) [91] | Deep CNN and gradient boosted tree | Breast cancer | Four classes and two of classification |
| Md Zahangir et al.(2018) [92] | IRRCNN(Inception Recurrent Residual CNN) | Breast cancer | Cancer classification and comparison with other models |
| M.M. Al Rahhal(2018) [93] | Deep CNN | Breast cancer | Detection and classification |
Table 3 represents different types of cancer detection and classification done by deep learning models using histopathological images. Mostly different CNN architectures are used for the wide area of cancer types such as lung, breast, brain, prostate, colon and colorectal cancer. This indicates the broad applicability of deep learning models in the various fields of cancer. Apart from these, deep learning can be applied to almost all areas. Some example applications are, object localization, object detection, image or video captioning [94], media and entertainment, image or video segmentation, autonomous car [95, 96], machine translation, speech recognition [97], security and defence [98, 99], vehicle classification [15], medicine and biology. There are also other challenging problems that have been resolved during the last few years, which could not be efficiently solved before this DL revolution. There are some recently conducted surveys in this area [100-102]. These documents investigate deep learning and its revolution.
6. DISCUSSION
In this study, the usefulness of machine learning and deep learning algorithms in different histopathological image analysis tasks, especially in oral cancer, was observed. The machine learning and deep learning algorithms were compared with diverse concerns in section 4. After studying the last nine years (2010-2018) of histopathological image analysis research work in computer vision for detecting various cancers, it was realized from the finding that the outcome of deep learning networks performs better compared to other traditional machine learning algorithms for handling complex queries. Within this period, there has been a rapid increase in several research studies using artificial intelligence techniques. Considering the number of layers, hierarchies, and concepts processed by the deep learning models, they are only suitable for carrying out complex calculations rather than simpler ones. The quality of data fed to the deep learning model determines the quality of the result. Because of the nested layers in the deep neural networks, these models do not require human intervention. Data are put through different layers of hierarchies and they can learn by their own errors, thus reducing the explicit intervention on the data engineering(pre-processing) process. In Table 1, although the findings indicated a very excellent efficiency in the prediction of oral cancer, they used very less number of samples (in the range of 10 to 110) for their study. Therefore, in the case of machine learning, an increase in the number of study samples is expected to affect the outcome and reduce the efficiency of the model and human intervention is required to retrain the model if the actual outcome is not the desired one.
A great deal of effort is currently being made to find effective methods to carry out unsupervised learning, as large quantities of unlabeled information are now becoming less costly, economically and technologically. Achievement in this goal will enable algorithms to know how the world operates by just watching it as we humans do.
Furthermore, real-world issues that generally involve high-dimensional ongoing state spaces (large number of states and/or actions) can make the issue intractable with present methods, severely restricting actual application growth. As an unsolved challenge, an effective way to cope with these kinds of issues continues.
Due to the capability of deep learning to execute feature engineering on its own, data scientists can save months of work for extracting the features from raw data. In deep learning, the data are scanned to look for features that correlate and combine them to allow faster learning without being specifically told to.
6.1. Challenges of Deep Learning
Deep learning approaches often face several challenges, starting with their own theoretical knowledge. An example of this is the lack of knowledge about the structure of the objective function in deep neural networks or why some architectures perform better than others.
There is a general absence of publicly accessible information in medical image analysis, and rare information labeled with high quality. Fewer than 100 patients are involved in the majority of the data sets described in this oral cancer evaluation. The articles in this survey report give good performance in the different assignments despite the small training data sets. Cho et al. [103] explained how many images are needed to train for medical image analysis problems. As a rule, data scientists say that “the more powerful abstraction you want, the more data is required”.
In medical image analysis, less training data or class inequality in the training set is also a major issue [104]. If appropriate quantities of training samples are not used, this may result in overfitting. Data inequality can be enhanced by using data augmentation [56, 64] to produce extra unusual image training samples, although there is a risk of overfitting.
In image recognition tasks, AI algorithms have outper- formed human execution and can deliver superior performance in medical image analysis to individuals. However, the issue occurs when a patient is misdiagnosed due to AI or AI-helped restorative administration or endures hopelessness. This is complemented by the failure to completely clarify how the black box of deep learning algorithms works.
6.2. Future Direction
In the section of literature review, the traditional machine learning applications for medical image analysis have been discussed.
The upcoming research areas include prognostication [105], content-based image retrieval [106, 107], caption generation [108, 109], and manipulation of physical objects with LSTMs and reinforcement learning [110, 111], involving surgical robots [112, 113].
These models highlight how the field of AI is quickly changing in medical image analysis and that numerous apps that have not yet been established may still exist. Thus, AI research area is very wide and needs further investigation. Further research is required in the following areas.
(1) Focusing more on adaptive dynamic programming (ADP) could produce a remarkable finding that accounts for more important brain research and computational intelligence.
(2) It would be interesting to construct deep learning models that can learn from fewer training data to make the prediction.
(3) Parameter tuning in AI is an emerging topic. Therefore, the effective use of the network variable’s optimization algorithms should be well studied.
(4) Research on the opportunities of integrating neural networks with other techniques that exist or develop.
(5) Hybrid neural network models should be developed for better efficiency and effectiveness.
CONCLUSION
Applications of different machine learning, as well as deep learning techniques highlighting the role of neural networks, are briefly discussed in this paper. Many methods are studied to classify images into distinct phases of cancer.
For getting enhanced performance, the use of increased dataset, selection, and extraction of appropriate features and combination of specified techniques may give adequate results. The proficiency of SVM is proved to be significant for different types of features showing its robustness in multiple fields. But it is a machine learning approach in which handcrafted features are fed to the model and works well for less number of the dataset. Mostly, CNN of deep learning works in an ideal way for image problems by doing the simultaneous work of feature extraction and classification. Implicit feature engineering and faster learning approach of CNN makes it possible to outperform in prediction /prognosis in the cancer domain. Finally, it can be said that CNN can work with a very large and different types of datasets to obtain closer to perfect results.
A more extensive review of deep learning methods within histopathology can be carried out for future work.
CONSENT FOR PUBLICATION
Not applicable.
FUNDING
None.
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
The authors thank reviewers of this work for their valuable comments and suggestions that improve the presentation of this research effort.


